pH-dependent conformation, dynamics, and aromatic interaction of the gating tryptophan residue of the influenza M2 proton channel from solid-state NMR
- PMID: 23601317
- PMCID: PMC3627873
- DOI: 10.1016/j.bpj.2013.02.054
pH-dependent conformation, dynamics, and aromatic interaction of the gating tryptophan residue of the influenza M2 proton channel from solid-state NMR
Abstract
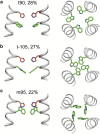
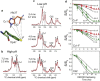
The M2 protein of the influenza virus conducts protons into the virion under external acidic pH. The proton selectivity of the tetrameric channel is controlled by a single histidine (His(37)), whereas channel gating is accomplished by a single tryptophan (Trp(41)) in the transmembrane domain of the protein. Aromatic interaction between these two functional residues has been previously observed in Raman spectra, but atomic-resolution evidence for this interaction remains scarce. Here we use high-resolution solid-state NMR spectroscopy to determine the side-chain conformation and dynamics of Trp(41) in the M2 transmembrane peptide by measuring the Trp chemical shifts, His(37)-Trp(41) distances, and indole dynamics at high and low pH. The interatomic distances constrain the Trp41 side-chain conformation to trans for χ1 and 120-135° for χ2. This t90 rotamer points the Nε1-Cε2-Cζ2 side of the indole toward the aqueous pore. The precise χ1 and χ2 angles differ by ∼20° between high and low pH. These differences, together with the known changes in the helix tilt angle between high and low pH, push the imidazole and indole rings closer together at low pH. Moreover, the measured order parameters indicate that the indole rings undergo simultaneous χ1 and χ2 torsional fluctuations at acidic pH, but only restricted χ1 fluctuations at high pH. As a result, the Trp(41) side chain periodically experiences strong cation-π interactions with His(37) at low pH as the indole sweeps through its trajectory, whereas at high pH the indole ring is further away from the imidazole. These results provide the structural basis for understanding how the His(37)-water proton exchange rate measured by NMR is reduced to the small proton flux measured in biochemical experiments. The indole dynamics, together with the known motion of the imidazolium, indicate that this compact ion channel uses economical side-chain dynamics to regulate proton conduction and gating.
Copyright © 2013 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures





Comment in
-
The functional heart of the M2 channel.Biophys J. 2013 Apr 16;104(8):1639-40. doi: 10.1016/j.bpj.2013.03.020. Biophys J. 2013. PMID: 23601309 Free PMC article. No abstract available.
Similar articles
-
Solid-state 19F NMR spectroscopy reveals that Trp41 participates in the gating mechanism of the M2 proton channel of influenza A virus.J Am Chem Soc. 2008 Jan 23;130(3):918-24. doi: 10.1021/ja0754305. Epub 2007 Dec 29. J Am Chem Soc. 2008. PMID: 18163621
-
Side-chain conformation of the M2 transmembrane peptide proton channel of influenza a virus from 19F solid-state NMR.J Phys Chem B. 2007 Sep 13;111(36):10825-32. doi: 10.1021/jp073823k. Epub 2007 Aug 17. J Phys Chem B. 2007. PMID: 17705425
-
Protonation of histidine and histidine-tryptophan interaction in the activation of the M2 ion channel from influenza a virus.Biochemistry. 2001 May 22;40(20):6053-60. doi: 10.1021/bi0028441. Biochemistry. 2001. PMID: 11352741
-
Roles of the histidine and tryptophan side chains in the M2 proton channel from influenza A virus.FEBS Lett. 2003 Sep 18;552(1):35-8. doi: 10.1016/s0014-5793(03)00781-6. FEBS Lett. 2003. PMID: 12972149 Review.
-
Structural basis for proton conduction and inhibition by the influenza M2 protein.Protein Sci. 2012 Nov;21(11):1620-33. doi: 10.1002/pro.2158. Epub 2012 Oct 9. Protein Sci. 2012. PMID: 23001990 Free PMC article. Review.
Cited by
-
Conformation and Trimer Association of the Transmembrane Domain of the Parainfluenza Virus Fusion Protein in Lipid Bilayers from Solid-State NMR: Insights into the Sequence Determinants of Trimer Structure and Fusion Activity.J Mol Biol. 2018 Mar 2;430(5):695-709. doi: 10.1016/j.jmb.2018.01.002. Epub 2018 Jan 10. J Mol Biol. 2018. PMID: 29330069 Free PMC article.
-
SARS-CoV-2 Envelope Protein Forms Clustered Pentamers in Lipid Bilayers.Biochemistry. 2022 Nov 1;61(21):2280-2294. doi: 10.1021/acs.biochem.2c00464. Epub 2022 Oct 11. Biochemistry. 2022. PMID: 36219675 Free PMC article.
-
Multiscale Simulation of an Influenza A M2 Channel Mutant Reveals Key Features of Its Markedly Different Proton Transport Behavior.J Am Chem Soc. 2022 Jan 19;144(2):769-776. doi: 10.1021/jacs.1c09281. Epub 2022 Jan 5. J Am Chem Soc. 2022. PMID: 34985907 Free PMC article.
-
Activation pH and Gating Dynamics of Influenza A M2 Proton Channel Revealed by Single-Molecule Spectroscopy.Angew Chem Int Ed Engl. 2017 May 2;56(19):5283-5287. doi: 10.1002/anie.201701874. Epub 2017 Apr 4. Angew Chem Int Ed Engl. 2017. PMID: 28374543 Free PMC article.
-
Fast Magic-Angle-Spinning 19F Spin Exchange NMR for Determining Nanometer 19F-19F Distances in Proteins and Pharmaceutical Compounds.J Phys Chem B. 2018 Mar 22;122(11):2900-2911. doi: 10.1021/acs.jpcb.8b00310. Epub 2018 Mar 13. J Phys Chem B. 2018. PMID: 29486126 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

