Average conformations determined from PRE data provide high-resolution maps of transient tertiary interactions in disordered proteins
- PMID: 23601321
- PMCID: PMC3628557
- DOI: 10.1016/j.bpj.2013.02.019
Average conformations determined from PRE data provide high-resolution maps of transient tertiary interactions in disordered proteins
Abstract
In the last decade it has become evident that disordered states of proteins play important physiological and pathological roles and that the transient tertiary interactions often present in these systems can play a role in their biological activity. The structural characterization of such states has so far largely relied on ensemble representations, which in principle account for both their local and global structural features. However, these approaches are inherently of low resolution due to the large number of degrees of freedom of conformational ensembles and to the sparse nature of the experimental data used to determine them. Here, we overcome these limitations by showing that tertiary interactions in disordered states can be mapped at high resolution by fitting paramagnetic relaxation enhancement data to a small number of conformations, which can be as low as one. This result opens up the possibility of determining the topology of cooperatively collapsed and hidden folded states when these are present in the vast conformational landscape accessible to disordered states of proteins. As a first application, we study the long-range tertiary interactions of acid-unfolded apomyoglobin from experimentally measured paramagnetic relaxation enhancement data.
Copyright © 2013 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures


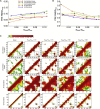

Comment in
-
Generating accurate contact maps of transient long-range interactions in intrinsically disordered proteins by paramagnetic relaxation enhancement.Biophys J. 2013 Apr 16;104(8):1635-6. doi: 10.1016/j.bpj.2013.01.060. Biophys J. 2013. PMID: 23601307 Free PMC article. No abstract available.
Similar articles
-
Generating accurate contact maps of transient long-range interactions in intrinsically disordered proteins by paramagnetic relaxation enhancement.Biophys J. 2013 Apr 16;104(8):1635-6. doi: 10.1016/j.bpj.2013.01.060. Biophys J. 2013. PMID: 23601307 Free PMC article. No abstract available.
-
Structural characterization of unfolded states of apomyoglobin using residual dipolar couplings.J Mol Biol. 2004 Jul 23;340(5):1131-42. doi: 10.1016/j.jmb.2004.05.022. J Mol Biol. 2004. PMID: 15236972
-
Structural interpretation of paramagnetic relaxation enhancement-derived distances for disordered protein states.J Mol Biol. 2009 Jul 17;390(3):467-77. doi: 10.1016/j.jmb.2009.05.019. Epub 2009 May 15. J Mol Biol. 2009. PMID: 19447112
-
Monitoring protein folding at atomic resolution.Acc Chem Res. 2004 Dec;37(12):929-36. doi: 10.1021/ar020156z. Acc Chem Res. 2004. PMID: 15609984 Review.
-
Intermediate states in protein folding.J Mol Biol. 1996 May 24;258(5):707-25. doi: 10.1006/jmbi.1996.0280. J Mol Biol. 1996. PMID: 8637003 Review.
Cited by
-
Concomitant disorder and high-affinity zinc binding in the human zinc- and iron-regulated transport protein 4 intracellular loop.Protein Sci. 2019 May;28(5):868-880. doi: 10.1002/pro.3591. Epub 2019 Mar 12. Protein Sci. 2019. PMID: 30793391 Free PMC article.
-
Generating accurate contact maps of transient long-range interactions in intrinsically disordered proteins by paramagnetic relaxation enhancement.Biophys J. 2013 Apr 16;104(8):1635-6. doi: 10.1016/j.bpj.2013.01.060. Biophys J. 2013. PMID: 23601307 Free PMC article. No abstract available.
-
Single residue modulators of amyloid formation in the N-terminal P1-region of α-synuclein.Nat Commun. 2022 Aug 25;13(1):4986. doi: 10.1038/s41467-022-32687-1. Nat Commun. 2022. PMID: 36008493 Free PMC article.
-
Visualizing transient dark states by NMR spectroscopy.Q Rev Biophys. 2015 Feb;48(1):35-116. doi: 10.1017/S0033583514000122. Q Rev Biophys. 2015. PMID: 25710841 Free PMC article.
-
PolyUbiquitin chain linkage topology selects the functions from the underlying binding landscape.PLoS Comput Biol. 2014 Jul 3;10(7):e1003691. doi: 10.1371/journal.pcbi.1003691. eCollection 2014 Jul. PLoS Comput Biol. 2014. PMID: 24992446 Free PMC article.
References
-
- Shortle D. The denatured state (the other half of the folding equation) and its role in protein stability. FASEB J. 1996;10:27–34. - PubMed
-
- Dobson C.M. Protein folding and misfolding. Nature. 2003;426:884–890. - PubMed
-
- Dunker A.K., Silman I., Sussman J.L. Function and structure of inherently disordered proteins. Curr. Opin. Struct. Biol. 2008;18:756–764. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

