A nucleus-targeted alternately spliced Nix/Bnip3L protein isoform modifies nuclear factor κB (NFκB)-mediated cardiac transcription
- PMID: 23603904
- PMCID: PMC3668707
- DOI: 10.1074/jbc.M113.452342
A nucleus-targeted alternately spliced Nix/Bnip3L protein isoform modifies nuclear factor κB (NFκB)-mediated cardiac transcription
Abstract
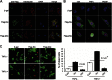
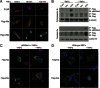
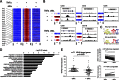
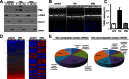
Several Bcl2 family proteins are expressed both as mitochondrial-targeted full-length and as cytosolic truncated alternately spliced isoforms. Recombinantly expressed shorter Bcl2 family isoforms can heterotypically bind to and prevent mitochondrial localization of their full-length analogs, thus suppressing their activity by sequestration. This "sponge" role requires 1:1 expression stoichiometry; absent this an alternate role is suggested. Here, RNA sequencing revealed coordinate regulation of BH3-only protein Nix/Bnip3L (Nix) and its alternately spliced soluble form (sNix) in hearts, but relative sNix/Nix expression of ∼1:10. Accordingly, we examined other putative functions of sNix. Although Nix expressed in H9c2 rat myoblasts localized to mitochondria, sNix showed variable cytoplasmic and nuclear distribution. Tumor necrosis factor α (TNFα) induced rapid and complete sNix nucleoplasmic translocation concomitant with nuclear translocation of the p65/RelA subunit of NFκB. sNix co-localized and co-precipitated with p65/RelA after TNFα stimulation; TNFα-induced sNix nuclear translocation did not occur in p65/RelA null murine embryonic fibroblasts. ChIP sequencing of TNFα-stimulated H9c2 cells revealed sNix suppression of p65/RelA binding to a subset of weaker DNA binding sites, accounting for its ability to alter gene expression in cultured cells and in vivo mouse hearts. These findings reveal TNFα-stimulated cytoplasmic-nuclear shuttling of the alternately spliced non-mitochondrial Nix isoform and uncover a role for sNix as a modulator of TNFα/NFκB-stimulated cardiac gene expression. Transcriptional co-regulation of sNix and Nix, combined with sNix posttranslational regulation by TNFα, comprises a previously unknown mechanism for molecular cross-talk between extrinsic death receptor and intrinsic mitochondrial apoptosis pathways.
Keywords: Bcl-2 Family Proteins; Cardiovascular Disease; Gene Regulation; NF-kappa B (NF-KB); Tumor Necrosis Factor (TNF); sNix.
Figures


Similar articles
-
Rad GTPase inhibits the NFκB pathway through interacting with RelA/p65 to impede its DNA binding and target gene transactivation.Cell Signal. 2014 Jul;26(7):1437-44. doi: 10.1016/j.cellsig.2014.03.003. Epub 2014 Mar 13. Cell Signal. 2014. PMID: 24632303
-
Mitochondrial death protein Nix is induced in cardiac hypertrophy and triggers apoptotic cardiomyopathy.Nat Med. 2002 Jul;8(7):725-30. doi: 10.1038/nm719. Epub 2002 Jun 10. Nat Med. 2002. PMID: 12053174
-
Tristetraprolin impairs NF-kappaB/p65 nuclear translocation.J Biol Chem. 2009 Oct 23;284(43):29571-81. doi: 10.1074/jbc.M109.031237. Epub 2009 Aug 4. J Biol Chem. 2009. PMID: 19654331 Free PMC article.
-
A brief overview of BNIP3L/NIX receptor-mediated mitophagy.FEBS Open Bio. 2021 Dec;11(12):3230-3236. doi: 10.1002/2211-5463.13307. Epub 2021 Oct 11. FEBS Open Bio. 2021. PMID: 34597467 Free PMC article. Review.
-
Nix Nought Nothing: fairy tale or real deal.J Mol Cell Cardiol. 2011 Oct;51(4):497-500. doi: 10.1016/j.yjmcc.2010.09.011. Epub 2010 Sep 18. J Mol Cell Cardiol. 2011. PMID: 20858501 Free PMC article. Review.
Cited by
-
LINC00365-SCGB2A1 axis inhibits the viability of breast cancer through targeting NF-κB signaling.Oncol Lett. 2020 Jan;19(1):753-762. doi: 10.3892/ol.2019.11166. Epub 2019 Nov 29. Oncol Lett. 2020. PMID: 31897191 Free PMC article.
-
Epigenetic coordination of embryonic heart transcription by dynamically regulated long noncoding RNAs.Proc Natl Acad Sci U S A. 2014 Aug 19;111(33):12264-9. doi: 10.1073/pnas.1410622111. Epub 2014 Jul 28. Proc Natl Acad Sci U S A. 2014. PMID: 25071214 Free PMC article.
-
Elimination of damaged mitochondria during UVB-induced senescence is orchestrated by NIX-dependent mitophagy.Aging Cell. 2024 Aug;23(8):e14186. doi: 10.1111/acel.14186. Epub 2024 May 17. Aging Cell. 2024. PMID: 38761001 Free PMC article.
-
Misoprostol regulates Bnip3 repression and alternative splicing to control cellular calcium homeostasis during hypoxic stress.Cell Death Discov. 2018 Sep 21;4:37. doi: 10.1038/s41420-018-0104-z. eCollection 2018. Cell Death Discov. 2018. PMID: 30275982 Free PMC article.
-
TUFT1 is expressed in breast cancer and involved in cancer cell proliferation and survival.Oncotarget. 2017 Aug 24;8(43):74962-74974. doi: 10.18632/oncotarget.20472. eCollection 2017 Sep 26. Oncotarget. 2017. PMID: 29088838 Free PMC article.
References
-
- Whelan R. S., Kaplinskiy V., Kitsis R. N. (2010) Cell death in the pathogenesis of heart disease: mechanisms and significance. Annu. Rev. Physiol. 72, 19–44 - PubMed
-
- Yussman M. G., Toyokawa T., Odley A., Lynch R. A., Wu G., Colbert M. C., Aronow B. J., Lorenz J. N., Dorn G. W., 2nd (2002) Mitochondrial death protein Nix is induced in cardiac hypertrophy and triggers apoptotic cardiomyopathy. Nat. Med. 8, 725–730 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

