Ubiquitin-specific protease 7 is a regulator of ubiquitin-conjugating enzyme UbE2E1
- PMID: 23603909
- PMCID: PMC3675629
- DOI: 10.1074/jbc.M113.469262
Ubiquitin-specific protease 7 is a regulator of ubiquitin-conjugating enzyme UbE2E1
Abstract
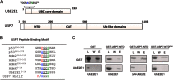
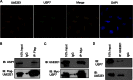
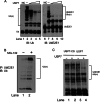
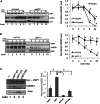
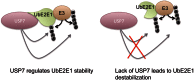
Ubiquitin-specific protease 7 (USP7) is a deubiquitinating enzyme found in all eukaryotes that catalyzes the removal of ubiquitin from specific target proteins. Here, we report that UbE2E1, an E2 ubiquitin conjugation enzyme with a unique N-terminal extension, is a novel USP7-interacting protein. USP7 forms a complex with UbE2E1 in vitro and in vivo through the ASTS USP7 binding motif within its N-terminal extension in an identical manner with other known USP7 binding proteins. We show that USP7 attenuates UbE2E1-mediated ubiquitination, an effect that requires the N-terminal ASTS sequence of UbE2E1 as well as the catalytic activity of USP7. Additionally, USP7 is critical in maintaining the steady state levels of UbE2E1 in cells. This study reveals a new cellular mechanism that couples the opposing activities of the ubiquitination machinery and a deubiquitinating enzyme to maintain and modulate the dynamic balance of the ubiquitin-proteasome system.
Keywords: Deubiquitination; E3 Ubiquitin Ligase; Protein Structure; Ubiquitin-conjugating Enzyme (Ubc); Ubiquitination.
Figures


Similar articles
-
UbE2E1/UBCH6 Is a Critical in Vivo E2 for the PRC1-catalyzed Ubiquitination of H2A at Lys-119.J Biol Chem. 2017 Feb 17;292(7):2893-2902. doi: 10.1074/jbc.M116.749564. Epub 2017 Jan 10. J Biol Chem. 2017. PMID: 28073915 Free PMC article.
-
OTUB1 non-catalytically stabilizes the E2 ubiquitin-conjugating enzyme UBE2E1 by preventing its autoubiquitination.J Biol Chem. 2018 Nov 23;293(47):18285-18295. doi: 10.1074/jbc.RA118.004677. Epub 2018 Oct 3. J Biol Chem. 2018. PMID: 30282802 Free PMC article.
-
Ubiquitin-specific protease 7 regulates nucleotide excision repair through deubiquitinating XPC protein and preventing XPC protein from undergoing ultraviolet light-induced and VCP/p97 protein-regulated proteolysis.J Biol Chem. 2014 Sep 26;289(39):27278-27289. doi: 10.1074/jbc.M114.589812. Epub 2014 Aug 12. J Biol Chem. 2014. PMID: 25118285 Free PMC article.
-
Current and future directions of USP7 interactome in cancer study.Biochim Biophys Acta Rev Cancer. 2023 Nov;1878(6):188992. doi: 10.1016/j.bbcan.2023.188992. Epub 2023 Sep 27. Biochim Biophys Acta Rev Cancer. 2023. PMID: 37775071 Review.
-
Regulation of USP7: A High Incidence of E3 Complexes.J Mol Biol. 2017 Nov 10;429(22):3395-3408. doi: 10.1016/j.jmb.2017.05.028. Epub 2017 Jun 4. J Mol Biol. 2017. PMID: 28591556 Review.
Cited by
-
USP7 inhibits Wnt/β-catenin signaling through promoting stabilization of Axin.Nat Commun. 2019 Sep 13;10(1):4184. doi: 10.1038/s41467-019-12143-3. Nat Commun. 2019. PMID: 31519875 Free PMC article.
-
USP7: Structure, substrate specificity, and inhibition.DNA Repair (Amst). 2019 Apr;76:30-39. doi: 10.1016/j.dnarep.2019.02.005. Epub 2019 Feb 16. DNA Repair (Amst). 2019. PMID: 30807924 Free PMC article. Review.
-
The emerging role of deubiquitinating enzymes in genomic integrity, diseases, and therapeutics.Cell Biosci. 2016 Dec 20;6:62. doi: 10.1186/s13578-016-0127-1. eCollection 2016. Cell Biosci. 2016. PMID: 28031783 Free PMC article. Review.
-
Identification and Characterization of USP7 Targets in Cancer Cells.Sci Rep. 2018 Oct 26;8(1):15833. doi: 10.1038/s41598-018-34197-x. Sci Rep. 2018. PMID: 30367141 Free PMC article.
-
HAUSP Is a Key Epigenetic Regulator of the Chromatin Effector Proteins.Genes (Basel). 2021 Dec 24;13(1):42. doi: 10.3390/genes13010042. Genes (Basel). 2021. PMID: 35052383 Free PMC article. Review.
References
-
- Pickart C. M. (2001) Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 - PubMed
-
- Schwartz A. L., Ciechanover A. (2009) Targeting proteins for destruction by the ubiquitin system: implications for human pathobiology. Annu. Rev. Pharmacol. Toxicol. 49, 73–96 - PubMed
-
- Jin J., Li X., Gygi S. P., Harper J. W. (2007) Dual E1 activation systems for ubiquitin differentially regulate E2 enzyme charging. Nature 447, 1135–1138 - PubMed
-
- van Wijk S. J., Timmers H. T. (2010) The family of ubiquitin-conjugating enzymes (E2s): deciding between life and death of proteins. FASEB J. 24, 981–993 - PubMed
-
- Deshaies R. J., Joazeiro C. A. (2009) RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

