Solution NMR refinement of a metal ion bound protein using metal ion inclusive restrained molecular dynamics methods
- PMID: 23609042
- PMCID: PMC3773525
- DOI: 10.1007/s10858-013-9729-7
Solution NMR refinement of a metal ion bound protein using metal ion inclusive restrained molecular dynamics methods
Abstract
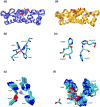
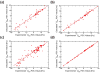
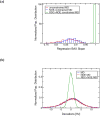
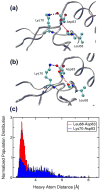
Correctly calculating the structure of metal coordination sites in a protein during the process of nuclear magnetic resonance (NMR) structure determination and refinement continues to be a challenging task. In this study, we present an accurate and convenient means by which to include metal ions in the NMR structure determination process using molecular dynamics (MD) simulations constrained by NMR-derived data to obtain a realistic and physically viable description of the metal binding site(s). This method provides the framework to accurately portray the metal ions and its binding residues in a pseudo-bond or dummy-cation like approach, and is validated by quantum mechanical/molecular mechanical (QM/MM) MD calculations constrained by NMR-derived data. To illustrate this approach, we refine the zinc coordination complex structure of the zinc sensing transcriptional repressor protein Staphylococcus aureus CzrA, generating over 130 ns of MD and QM/MM MD NMR-data compliant sampling. In addition to refining the first coordination shell structure of the Zn(II) ion, this protocol benefits from being performed in a periodically replicated solvation environment including long-range electrostatics. We determine that unrestrained (not based on NMR data) MD simulations correlated to the NMR data in a time-averaged ensemble. The accurate solution structure ensemble of the metal-bound protein accurately describes the role of conformational sampling in allosteric regulation of DNA binding by zinc and serves to validate our previous unrestrained MD simulations of CzrA. This methodology has potentially broad applicability in the structure determination of metal ion bound proteins, protein folding and metal template protein-design studies.
Figures

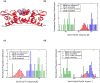
References
-
- Allen MP, Tildesley DJ. Computer Simulations of Liquids. Clarendon Press; Oxford: 1987.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

