A multi-layer inference approach to reconstruct condition-specific genes and their regulation
- PMID: 23610368
- PMCID: PMC3673221
- DOI: 10.1093/bioinformatics/btt186
A multi-layer inference approach to reconstruct condition-specific genes and their regulation
Abstract
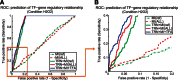
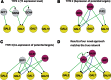
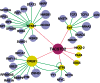
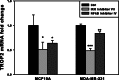
An important topic in systems biology is the reverse engineering of regulatory mechanisms through reconstruction of context-dependent gene networks. A major challenge is to identify the genes and the regulations specific to a condition or phenotype, given that regulatory processes are highly connected such that a specific response is typically accompanied by numerous collateral effects. In this study, we design a multi-layer approach that is able to reconstruct condition-specific genes and their regulation through an integrative analysis of large-scale information of gene expression, protein interaction and transcriptional regulation (transcription factor-target gene relationships). We establish the accuracy of our methodology against synthetic datasets, as well as a yeast dataset. We then extend the framework to the application of higher eukaryotic systems, including human breast cancer and Arabidopsis thaliana cold acclimation. Our study identified TACSTD2 (TROP2) as a target gene for human breast cancer and discovered its regulation by transcription factors CREB, as well as NFkB. We also predict KIF2C is a target gene for ER-/HER2- breast cancer and is positively regulated by E2F1. The predictions were further confirmed through experimental studies.
Availability: The implementation and detailed protocol of the layer approach is available at http://www.egr.msu.edu/changroup/Protocols/Three-layer%20approach%20 to % 20reconstruct%20condition.html.
Figures


Similar articles
-
Rapid transcriptional and metabolic regulation of the deacclimation process in cold acclimated Arabidopsis thaliana.BMC Genomics. 2017 Sep 16;18(1):731. doi: 10.1186/s12864-017-4126-3. BMC Genomics. 2017. PMID: 28915789 Free PMC article.
-
Identification of novel targets for breast cancer by exploring gene switches on a genome scale.BMC Genomics. 2011 Nov 3;12:547. doi: 10.1186/1471-2164-12-547. BMC Genomics. 2011. PMID: 22053771 Free PMC article.
-
Identification of small RNAs during cold acclimation in Arabidopsis thaliana.BMC Plant Biol. 2020 Jun 29;20(1):298. doi: 10.1186/s12870-020-02511-3. BMC Plant Biol. 2020. PMID: 32600430 Free PMC article.
-
Gene Regulatory Networks Mediating Cold Acclimation: The CBF Pathway.Adv Exp Med Biol. 2018;1081:3-22. doi: 10.1007/978-981-13-1244-1_1. Adv Exp Med Biol. 2018. PMID: 30288701 Review.
-
Computational and experimental approaches for modeling gene regulatory networks.Curr Pharm Des. 2007;13(14):1415-36. doi: 10.2174/138161207780765945. Curr Pharm Des. 2007. PMID: 17504165 Review.
Cited by
-
TACSTD2 upregulation is an early reaction to lung infection.Sci Rep. 2022 Jun 10;12(1):9583. doi: 10.1038/s41598-022-13637-9. Sci Rep. 2022. PMID: 35688908 Free PMC article.
-
Trop2: Jack of All Trades, Master of None.Cancers (Basel). 2020 Nov 11;12(11):3328. doi: 10.3390/cancers12113328. Cancers (Basel). 2020. PMID: 33187148 Free PMC article. Review.
-
Trop2 and its overexpression in cancers: regulation and clinical/therapeutic implications.Genes Cancer. 2015 Mar;6(3-4):84-105. doi: 10.18632/genesandcancer.40. Genes Cancer. 2015. PMID: 26000093 Free PMC article. Review.
-
Design and analysis of a Petri net model of the Von Hippel-Lindau (VHL) tumor suppressor interaction network.PLoS One. 2014 Jun 2;9(6):e96986. doi: 10.1371/journal.pone.0096986. eCollection 2014. PLoS One. 2014. PMID: 24886840 Free PMC article.
-
KIF2C/MCAK a prognostic biomarker and its oncogenic potential in malignant progression, and prognosis of cancer patients: a systematic review and meta-analysis as biomarker.Crit Rev Clin Lab Sci. 2024 Sep;61(6):404-434. doi: 10.1080/10408363.2024.2309933. Epub 2024 Feb 12. Crit Rev Clin Lab Sci. 2024. PMID: 38344808 Free PMC article.
References
-
- Abdel-Fatah T, et al. P4-09-11: kinesin family member 2C (KIF2C) is a new surrogate prognostic marker in breast cancer (BC) Cancer Res. 2012;71:P4–09–11.
-
- Agarwal M, et al. A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J. Biol. Chem. 2006;281:37636–37645. - PubMed
-
- Almuallim H, Dietterich TG. Proceedings of The Ninth National Conference on Artificial Intelligence. Menlo Park, CA: AAAI Press; 1991. Learning with many irrelevant features; pp. 547–552.
-
- Basso K, et al. Reverse engineering of regulatory networks in human B cells. Nat. Genet. 2005;37:382–390. - PubMed
-
- Bontempi G, Meyer PE. Causal filter selection in microarray data. ICML. 2010:95–102.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

