Escherichia coli rimM and yjeQ null strains accumulate immature 30S subunits of similar structure and protein complement
- PMID: 23611982
- PMCID: PMC3683913
- DOI: 10.1261/rna.037523.112
Escherichia coli rimM and yjeQ null strains accumulate immature 30S subunits of similar structure and protein complement
Abstract
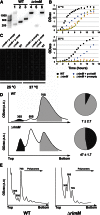
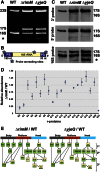
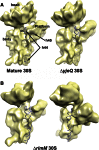
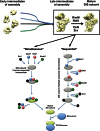
Assembly of the Escherichia coli 30S ribosomal subunits proceeds through multiple parallel pathways. The protein factors RimM, YjeQ, RbfA, and Era work in conjunction to assist at the late stages of the maturation process of the small subunit. However, it is unclear how the functional interplay between these factors occurs in the context of multiple parallel pathways. To understand how these factors work together, we have characterized the immature 30S subunits that accumulate in ΔrimM cells and compared them with immature 30S subunits from a ΔyjeQ strain. The cryo-EM maps obtained from these particles showed that the densities representing helices 44 and 45 in the rRNA were partially missing, suggesting mobility of these motifs. These 30S subunits were also partially depleted in all tertiary ribosomal proteins, particularly those binding in the head domain. Using image classification, we identified four subpopulations of ΔrimM immature 30S subunits differing in the amount of missing density for helices 44 and 45, as well as the amount of density existing in these maps for the underrepresented proteins. The structural defects found in these immature subunits resembled those of the 30S subunits that accumulate in the ΔyjeQ strain. These findings are consistent with an "early convergency model" in which multiple parallel assembly pathways of the 30S subunit converge into a late assembly intermediate, as opposed to the mature state. Functionally related factors will bind to this intermediate to catalyze the last steps of maturation leading to the mature 30S subunit.
Keywords: 30S subunit; RimM protein; YjeQ protein; cryo-electron microscopy; ribosome assembly.
Figures
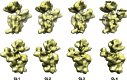

Similar articles
-
The C-terminal helix in the YjeQ zinc-finger domain catalyzes the release of RbfA during 30S ribosome subunit assembly.RNA. 2015 Jun;21(6):1203-16. doi: 10.1261/rna.049171.114. Epub 2015 Apr 22. RNA. 2015. PMID: 25904134 Free PMC article.
-
Binding properties of YjeQ (RsgA), RbfA, RimM and Era to assembly intermediates of the 30S subunit.Nucleic Acids Res. 2016 Nov 16;44(20):9918-9932. doi: 10.1093/nar/gkw613. Epub 2016 Jul 5. Nucleic Acids Res. 2016. PMID: 27382067 Free PMC article.
-
Role of Era in assembly and homeostasis of the ribosomal small subunit.Nucleic Acids Res. 2019 Sep 5;47(15):8301-8317. doi: 10.1093/nar/gkz571. Nucleic Acids Res. 2019. PMID: 31265110 Free PMC article.
-
Assembly of bacterial ribosomes.Annu Rev Biochem. 2011;80:501-26. doi: 10.1146/annurev-biochem-062608-160432. Annu Rev Biochem. 2011. PMID: 21529161 Review.
-
Structural insights into cell cycle control by essential GTPase Era.Postepy Biochem. 2016;62(3):335-342. Postepy Biochem. 2016. PMID: 28132488 Free PMC article. Review.
Cited by
-
Emerging Quantitative Biochemical, Structural, and Biophysical Methods for Studying Ribosome and Protein-RNA Complex Assembly.Biomolecules. 2023 May 19;13(5):866. doi: 10.3390/biom13050866. Biomolecules. 2023. PMID: 37238735 Free PMC article. Review.
-
A conserved rRNA switch is central to decoding site maturation on the small ribosomal subunit.Sci Adv. 2021 Jun 4;7(23):eabf7547. doi: 10.1126/sciadv.abf7547. Print 2021 Jun. Sci Adv. 2021. PMID: 34088665 Free PMC article.
-
Alternative conformations and motions adopted by 30S ribosomal subunits visualized by cryo-electron microscopy.RNA. 2020 Dec;26(12):2017-2030. doi: 10.1261/rna.075846.120. Epub 2020 Sep 28. RNA. 2020. PMID: 32989043 Free PMC article.
-
Hfq and RNase R Mediate rRNA Processing and Degradation in a Novel RNA Quality Control Process.mBio. 2020 Oct 20;11(5):e02398-20. doi: 10.1128/mBio.02398-20. mBio. 2020. PMID: 33082262 Free PMC article.
-
The C-terminal helix in the YjeQ zinc-finger domain catalyzes the release of RbfA during 30S ribosome subunit assembly.RNA. 2015 Jun;21(6):1203-16. doi: 10.1261/rna.049171.114. Epub 2015 Apr 22. RNA. 2015. PMID: 25904134 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
