Site-specific acetylation of the proteasome activator REGγ directs its heptameric structure and functions
- PMID: 23612972
- PMCID: PMC3675592
- DOI: 10.1074/jbc.M112.437129
Site-specific acetylation of the proteasome activator REGγ directs its heptameric structure and functions
Abstract
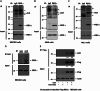
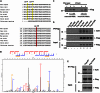
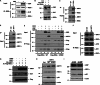
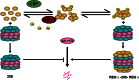
The proteasome activator REGγ has been reported to promote degradation of steroid receptor coactivator-3 and cyclin-dependent kinase inhibitors p21, p16, and p19 in a ubiquitin- and ATP-independent manner. A recent comparative analysis of REGγ expression in mouse and human tissues reveals a unique pattern of REGγ in specific cell types, suggesting undisclosed functions and biological importance of this molecule. Despite the emerging progress made in REGγ-related studies, how REGγ function is regulated remains to be explored. In this study, we report for the first time that REGγ can be acetylated mostly on its lysine 195 (Lys-195) residue by CREB binding protein (CBP), which can be reversed by sirtuin 1 (SIRT1) in mammalian cells. Site-directed mutagenesis abrogated acetylation at Lys-195 and significantly attenuated the capability of REGγ to degrade its target substrates, p21 and hepatitis C virus core protein. Mechanistically, acetylation at Lys-195 is important for the interactions between REGγ monomers and ultimately influences REGγ heptamerization. Biological analysis of cells containing REGγ-WT or REGγ-K195R mutant indicates an impact of acetylation on REGγ-mediated regulation of cell proliferation and cell cycle progression. These findings reveal a previously unknown mechanism in the regulation of REGγ assembly and activity, suggesting a potential venue for the intervention of the ubiquitin-independent REGγ proteasome activity.
Keywords: Acetylation; Cell Biology; Gene Regulation; Heptamerization; K195; Proteasome; Protein Degradation; Protein Self-assembly; Protein-Protein Interactions; REGγ.
Figures


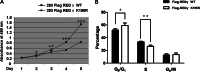
References
-
- Dubiel W., Pratt G., Ferrell K., Rechsteiner M. (1992) Purification of an 11 S regulator of the multicatalytic protease. J. Biol. Chem. 267, 22369–22377 - PubMed
-
- Ma C. P., Slaughter C. A., DeMartino G. N. (1992) Identification, purification, and characterization of a protein activator (PA28) of the 20 S proteasome (macropain). J. Biol. Chem. 267, 10515–10523 - PubMed
-
- Li X., Amazit L., Long W., Lonard D. M., Monaco J. J., O'Malley B. W. (2007) Ubiquitin- and ATP-independent proteolytic turnover of p21 by the REGγ-proteasome pathway. Mol. Cell 26, 831–842 - PubMed
-
- Li X., Lonard D. M., Jung S. Y., Malovannaya A., Feng Q., Qin J., Tsai S. Y., Tsai M. J., O'Malley B. W. (2006) The SRC-3/AIB1 coactivator is degraded in a ubiquitin- and ATP-independent manner by the REGγ proteasome. Cell 124, 381–392 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

