Bridge-induced chromosome translocation in yeast relies upon a Rad54/Rdh54-dependent, Pol32-independent pathway
- PMID: 23613757
- PMCID: PMC3629078
- DOI: 10.1371/journal.pone.0060926
Bridge-induced chromosome translocation in yeast relies upon a Rad54/Rdh54-dependent, Pol32-independent pathway
Abstract
While in mammalian cells the genetic determinism of chromosomal translocation remains unclear, the yeast Saccharomyces cerevisiae has become an ideal model system to generate ad hoc translocations and analyze their cellular and molecular outcome. A linear DNA cassette carrying a selectable marker flanked by perfect homologies to two chromosomes triggers a bridge-induced translocation (BIT) in budding yeast, with variable efficiency. A postulated two-step process to produce BIT translocants is based on the cooperation between the Homologous Recombination System (HRS) and Break-Induced Replication (BIR); however, a clear indication of the molecular factors underlying the genetic mechanism is still missing. In this work we provide evidence that BIT translocation is elicited by the Rad54 helicase and completed by a Pol32-independent replication pathway. Our results demonstrate also that Rdh54 is involved in the stability of the translocants, suggesting a mitotic role in chromosome pairing and segregation. Moreover, when RAD54 is over-expressed, an ensemble of secondary rearrangements between repeated DNA tracts arise after the initial translocation event, leading to severe aneuploidy with loss of genetic material, which prompts the identification of fragile sites within the yeast genome.
Conflict of interest statement
Figures
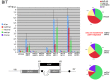

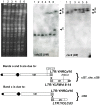
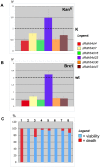
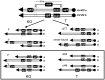
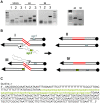

Similar articles
-
Chromosomal translocations caused by either pol32-dependent or pol32-independent triparental break-induced replication.Mol Cell Biol. 2009 Oct;29(20):5441-54. doi: 10.1128/MCB.00256-09. Epub 2009 Aug 3. Mol Cell Biol. 2009. PMID: 19651902 Free PMC article.
-
Rad54 and Rdh54 occupy spatially and functionally distinct sites within the Rad51-ssDNA presynaptic complex.EMBO J. 2020 Oct 15;39(20):e105705. doi: 10.15252/embj.2020105705. Epub 2020 Aug 13. EMBO J. 2020. PMID: 32790929 Free PMC article.
-
Rdh54/Tid1 inhibits Rad51-Rad54-mediated D-loop formation and limits D-loop length.Elife. 2020 Nov 13;9:e59112. doi: 10.7554/eLife.59112. Elife. 2020. PMID: 33185188 Free PMC article.
-
Discrete roles for Rad54 and Rdh54 during homologous recombination.Curr Opin Genet Dev. 2021 Dec;71:48-54. doi: 10.1016/j.gde.2021.06.013. Epub 2021 Jul 20. Curr Opin Genet Dev. 2021. PMID: 34293661 Review.
-
Investigation of Break-Induced Replication in Yeast.Methods Enzymol. 2018;601:161-203. doi: 10.1016/bs.mie.2017.12.010. Epub 2018 Feb 3. Methods Enzymol. 2018. PMID: 29523232 Review.
Cited by
-
The genome-wide rate and spectrum of spontaneous mutations differ between haploid and diploid yeast.Proc Natl Acad Sci U S A. 2018 May 29;115(22):E5046-E5055. doi: 10.1073/pnas.1801040115. Epub 2018 May 14. Proc Natl Acad Sci U S A. 2018. PMID: 29760081 Free PMC article.
-
Fluconazole induces ROS in Cryptococcus neoformans and contributes to DNA damage in vitro.PLoS One. 2018 Dec 7;13(12):e0208471. doi: 10.1371/journal.pone.0208471. eCollection 2018. PLoS One. 2018. PMID: 30532246 Free PMC article.
-
High reactive oxygen species levels are detected at the end of the chronological life span of translocant yeast cells.Mol Genet Genomics. 2016 Feb;291(1):423-35. doi: 10.1007/s00438-015-1120-9. Epub 2015 Sep 30. Mol Genet Genomics. 2016. PMID: 26423068
-
Per aspera ad astra: When harmful chromosomal translocations become a plus value in genetic evolution. Lessons from Saccharomyces cerevisiae.Microb Cell. 2015 Aug 20;2(10):363-375. doi: 10.15698/mic2015.10.230. Microb Cell. 2015. PMID: 28357264 Free PMC article. Review.
-
Bridge-Induced Translocation between NUP145 and TOP2 Yeast Genes Models the Genetic Fusion between the Human Orthologs Associated With Acute Myeloid Leukemia.Front Oncol. 2017 Sep 29;7:231. doi: 10.3389/fonc.2017.00231. eCollection 2017. Front Oncol. 2017. PMID: 29034209 Free PMC article.
References
-
- Poitras JL, Costa D, Kluk MJ, Amrein PC, Stone RM, et al. (2011) Genomic alterations in myeloid neoplasms with novel, apparently balanced translocations. Cancer Genet 204: 68–76. - PubMed
-
- Klemke M, Drieschner N, Laabs A, Rippe V, Belge G, et al. (2011) On the prevalence of the PAX8-PPARG fusion resulting from the chromosomal translocation t(2;3)(q13;p25) in adenomas of the thyroid. Cancer Genet 204: 334–339. - PubMed
-
- Hervé AL, Florence M, Philippe M, Michel A, Thierry F, et al. (2011) Molecular heterogeneity of multiple myeloma: pathogenesis, prognosis, and therapeutic implications. J Clin Oncol 29: 1893–1897. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

