Molecular dynamics simulations of depth distribution of spin-labeled phospholipids within lipid bilayer
- PMID: 23614631
- PMCID: PMC3713849
- DOI: 10.1021/jp4026706
Molecular dynamics simulations of depth distribution of spin-labeled phospholipids within lipid bilayer
Abstract
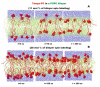
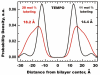
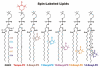
Spin-labeled lipids are commonly used as fluorescence quenchers in studies of membrane penetration of dye-labeled proteins and peptides using depth-dependent quenching. Accurate calculations of depth of the fluorophore rely on the use of several spin labels placed in the membrane at various positions. The depth of the quenchers (spin probes) has to be determined independently; however, experimental determination of transverse distributions of spin probe depths is difficult. In this Article, we use molecular dynamics (MD) simulations to study the membrane behavior and depth distributions of spin-labeled phospholipids in a 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) bilayer. To probe different depths within the bilayer, a series containing five Doxyl-labeled lipids (n-Doxyl PC) has been studied, in which a spin moiety was covalently attached to nth carbon atoms (where n = 5, 7, 10, 12, and 14) of the sn-2 stearoyl chain of the host phospholipid. Our results demonstrate that the chain-attached spin labels are broadly distributed across the model membrane and their environment is characterized by a high degree of mobility and structural heterogeneity. Despite the high thermal disorder, the depth distributions of the Doxyl labels were found to correlate well with their attachment positions, indicating that the distribution of the spin label within the model membrane is dictated by the depth of the nth lipid carbon atom and not by intrinsic properties of the label. In contrast, a much broader and heterogeneous distribution was observed for a headgroup-attached Tempo spin label of Tempo-PC lipids. MD simulations reveal that, due to the hydrophobic nature, a Tempo moiety favors partitioning from the headgroup region deeper into the membrane. Depending on the concentration of Tempo-PC lipids, the probable depth of the Tempo moiety could span a range from 14.4 to 18.2 Å from the membrane center. Comparison of the MD-estimated immersion depths of Tempo and n-Doxyl labels with their suggested experimental depth positions allows us to review critically the possible sources of error in depth-dependent fluorescence quenching studies.
Figures





References
-
- Klug CS, Feix JB. Methods and applications of site-directed spin labeling EPR spectroscopy. Methods in Cell Biology. 2008;Vol. 84:617–658. - PubMed
-
- Chattopadhyay A, London E. Parallax method for direct measurement of membrane penetration depth utilizing fluorescence quenching by spin-labeled phospholipids. Biochemistry. 1987;26(1):39–45. - PubMed
-
- Ladokhin AS. Distribution analysis of depth-dependent fluorescence quenching in membranes: A practical guide. In: Ludwig Brand MLJ, editor. Methods in Enzymology. Volume 278. Academic Press; 1997. pp. 462–473. - PubMed
-
- Ladokhin AS. Evaluation of lipid exposure of tryptophan residues in membrane peptides and proteins. Analytical Biochemistry. 1999;276(2):65–71. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

