Specified neural progenitors sort to form sharp domains after noisy Shh signaling
- PMID: 23622240
- PMCID: PMC3674856
- DOI: 10.1016/j.cell.2013.03.023
Specified neural progenitors sort to form sharp domains after noisy Shh signaling
Abstract
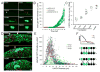
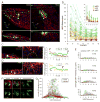
Sharply delineated domains of cell types arise in developing tissues under instruction of inductive signal (morphogen) gradients, which specify distinct cell fates at different signal levels. The translation of a morphogen gradient into discrete spatial domains relies on precise signal responses at stable cell positions. However, cells in developing tissues undergoing morphogenesis and proliferation often experience complex movements, which may affect their morphogen exposure, specification, and positioning. How is a clear pattern achieved with cells moving around? Using in toto imaging of the zebrafish neural tube, we analyzed specification patterns and movement trajectories of neural progenitors. We found that specified progenitors of different fates are spatially mixed following heterogeneous Sonic Hedgehog signaling responses. Cell sorting then rearranges them into sharply bordered domains. Ectopically induced motor neuron progenitors also robustly sort to correct locations. Our results reveal that cell sorting acts to correct imprecision of spatial patterning by noisy inductive signals.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures


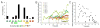


Comment in
-
Sorting sloppy Sonic.Cell. 2013 Apr 25;153(3):509-10. doi: 10.1016/j.cell.2013.04.021. Cell. 2013. PMID: 23622235 Free PMC article.
References
-
- Ahn S, Joyner AL. Dynamic changes in the response of cells to positive hedgehog signaling during mouse limb patterning. Cell. 2004;118:505–516. - PubMed
-
- Catala M, Teillet MA, De Robertis EM, Le Douarin ML. A spinal cord fate map in the avian embryo: while regressing, Hensen’s node lays down the notochord and floor plate thus joining the spinal cord lateral walls. Development. 1996;122:2599–2610. - PubMed
-
- Chamberlain CE, Jeong J, Guo C, Allen BL, McMahon AP. Notochord-derived Shh concentrates in close association with the apically positioned basal body in neural target cells and forms a dynamic gradient during neural patterning. Development. 2008;135:1097–1106. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

