14-3-3 proteins interact with a hybrid prenyl-phosphorylation motif to inhibit G proteins
- PMID: 23622247
- PMCID: PMC3690454
- DOI: 10.1016/j.cell.2013.03.044
14-3-3 proteins interact with a hybrid prenyl-phosphorylation motif to inhibit G proteins
Erratum in
- Cell. 2013 May 23;153(5):1164
Abstract
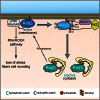
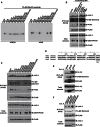
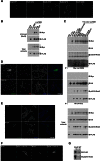
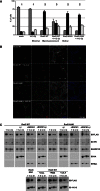
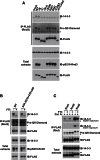
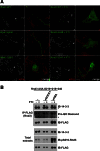
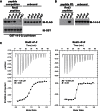
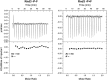
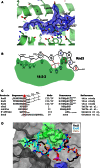
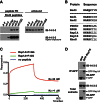
Signaling through G proteins normally involves conformational switching between GTP- and GDP-bound states. Several Rho GTPases are also regulated by RhoGDI binding and sequestering in the cytosol. Rnd proteins are atypical constitutively GTP-bound Rho proteins, whose regulation remains elusive. Here, we report a high-affinity 14-3-3-binding site at the C terminus of Rnd3 consisting of both the Cys241-farnesyl moiety and a Rho-associated coiled coil containing protein kinase (ROCK)-dependent Ser240 phosphorylation site. 14-3-3 binding to Rnd3 also involves phosphorylation of Ser218 by ROCK and/or Ser210 by protein kinase C (PKC). The crystal structure of a phosphorylated, farnesylated Rnd3 peptide with 14-3-3 reveals a hydrophobic groove in 14-3-3 proteins accommodating the farnesyl moiety. Functionally, 14-3-3 inhibits Rnd3-induced cell rounding by translocating it from the plasma membrane to the cytosol. Rnd1, Rnd2, and geranylgeranylated Rap1A interact similarly with 14-3-3. In contrast to the canonical GTP/GDP switch that regulates most Ras superfamily members, our results reveal an unprecedented mechanism for G protein inhibition by 14-3-3 proteins.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures




Comment in
-
Cell signalling: New G protein family rules.Nat Rev Mol Cell Biol. 2013 Jun;14(6):326. doi: 10.1038/nrm3590. Epub 2013 May 15. Nat Rev Mol Cell Biol. 2013. PMID: 23673967 No abstract available.
Similar articles
-
Interaction between Rho GTPases and 14-3-3 Proteins.Int J Mol Sci. 2017 Oct 15;18(10):2148. doi: 10.3390/ijms18102148. Int J Mol Sci. 2017. PMID: 29036929 Free PMC article. Review.
-
Rnd3-induced cell rounding requires interaction with Plexin-B2.J Cell Sci. 2016 Nov 1;129(21):4046-4056. doi: 10.1242/jcs.192211. Epub 2016 Sep 21. J Cell Sci. 2016. PMID: 27656111 Free PMC article.
-
The GTPase-deficient Rnd proteins are stabilized by their effectors.J Biol Chem. 2012 Sep 7;287(37):31311-20. doi: 10.1074/jbc.M111.327056. Epub 2012 Jul 17. J Biol Chem. 2012. PMID: 22807448 Free PMC article.
-
Regulation of Rnd3 localization and function by protein kinase C alpha-mediated phosphorylation.Biochem J. 2009 Oct 23;424(1):153-61. doi: 10.1042/BJ20082377. Biochem J. 2009. PMID: 19723022 Free PMC article.
-
Function and regulation of RhoE.Biochem Soc Trans. 2005 Aug;33(Pt 4):649-51. doi: 10.1042/BST0330649. Biochem Soc Trans. 2005. PMID: 16042565 Review.
Cited by
-
RhoA/mDia-1/profilin-1 signaling targets microvascular endothelial dysfunction in diabetic retinopathy.Graefes Arch Clin Exp Ophthalmol. 2015 May;253(5):669-80. doi: 10.1007/s00417-015-2985-3. Epub 2015 Mar 20. Graefes Arch Clin Exp Ophthalmol. 2015. PMID: 25791356 Review.
-
A RhoA and Rnd3 cycle regulates actin reassembly during membrane blebbing.Proc Natl Acad Sci U S A. 2016 Mar 29;113(13):E1863-71. doi: 10.1073/pnas.1600968113. Epub 2016 Mar 14. Proc Natl Acad Sci U S A. 2016. PMID: 26976596 Free PMC article.
-
Interaction between Rho GTPases and 14-3-3 Proteins.Int J Mol Sci. 2017 Oct 15;18(10):2148. doi: 10.3390/ijms18102148. Int J Mol Sci. 2017. PMID: 29036929 Free PMC article. Review.
-
Arsenite stress down-regulates phosphorylation and 14-3-3 binding of leucine-rich repeat kinase 2 (LRRK2), promoting self-association and cellular redistribution.J Biol Chem. 2014 Aug 1;289(31):21386-400. doi: 10.1074/jbc.M113.528463. Epub 2014 Jun 18. J Biol Chem. 2014. PMID: 24942733 Free PMC article.
-
The N- and C-terminal ends of RPGR can bind to PDE6δ.EMBO Rep. 2015 Dec;16(12):1583-5. doi: 10.15252/embr.201541404. Epub 2015 Nov 9. EMBO Rep. 2015. PMID: 26553937 Free PMC article.
References
-
- Bokoch G.M., Quilliam L.A., Bohl B.P., Jesaitis A.J., Quinn M.T. Inhibition of Rap1A binding to cytochrome b558 of NADPH oxidase by phosphorylation of Rap1A. Science. 1991;254:1794–1796. - PubMed
-
- Chandra A., Grecco H.E., Pisupati V., Perera D., Cassidy L., Skoulidis F., Ismail S.A., Hedberg C., Hanzal-Bayer M., Venkitaraman A.R. The GDI-like solubilizing factor PDEδ sustains the spatial organization and signalling of Ras family proteins. Nat. Cell Biol. 2012;14:148–158. - PubMed
Supplemental References
-
- Corey E., Takeda M., Kim C. Method for selective conversion of allylic and benzylic alcohols to halides under neutral conditions. Tetrahedron Lett. 1972;13:4339–4342.
-
- Dalal S.N., Yaffe M.B., DeCaprio J.A. 14-3-3 family members act coordinately to regulate mitotic progression. Cell Cycle. 2004;3:672–677. - PubMed
-
- O’Reilly N., Charbin A., Lopez-Serra L., Uhlmann F. Facile synthesis of budding yeast a-factor and its use to synchronize cells of α mating type. Yeast. 2012;29:233–240. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

