Practical use of chemical shift databases for protein solid-state NMR: 2D chemical shift maps and amino-acid assignment with secondary-structure information
- PMID: 23625364
- PMCID: PMC4048757
- DOI: 10.1007/s10858-013-9732-z
Practical use of chemical shift databases for protein solid-state NMR: 2D chemical shift maps and amino-acid assignment with secondary-structure information
Abstract
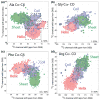
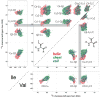
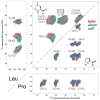
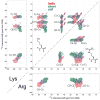
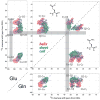
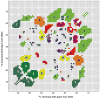
We introduce a Python-based program that utilizes the large database of (13)C and (15)N chemical shifts in the Biological Magnetic Resonance Bank to rapidly predict the amino acid type and secondary structure from correlated chemical shifts. The program, called PACSYlite Unified Query (PLUQ), is designed to help assign peaks obtained from 2D (13)C-(13)C, (15)N-(13)C, or 3D (15)N-(13)C-(13)C magic-angle-spinning correlation spectra. We show secondary-structure specific 2D (13)C-(13)C correlation maps of all twenty amino acids, constructed from a chemical shift database of 262,209 residues. The maps reveal interesting conformation-dependent chemical shift distributions and facilitate searching of correlation peaks during amino-acid type assignment. Based on these correlations, PLUQ outputs the most likely amino acid types and the associated secondary structures from inputs of experimental chemical shifts. We test the assignment accuracy using four high-quality protein structures. Based on only the Cα and Cβ chemical shifts, the highest-ranked PLUQ assignments were 40-60 % correct in both the amino-acid type and the secondary structure. For three input chemical shifts (CO-Cα-Cβ or N-Cα-Cβ), the first-ranked assignments were correct for 60 % of the residues, while within the top three predictions, the correct assignments were found for 80 % of the residues. PLUQ and the chemical shift maps are expected to be useful at the first stage of sequential assignment, for combination with automated sequential assignment programs, and for highly disordered proteins for which secondary structure analysis is the main goal of structure determination.
Figures

Similar articles
-
Automated amino acid side-chain NMR assignment of proteins using (13)C- and (15)N-resolved 3D [ (1)H, (1)H]-NOESY.J Biomol NMR. 2008 Sep;42(1):23-33. doi: 10.1007/s10858-008-9259-x. Epub 2008 Aug 16. J Biomol NMR. 2008. PMID: 18709333
-
Optimal-Control-Based Cβ Chemical Shift Encoding for Efficient Signal Assignment of Solid Proteins.J Phys Chem B. 2023 Nov 30;127(47):10118-10128. doi: 10.1021/acs.jpcb.3c05914. Epub 2023 Nov 17. J Phys Chem B. 2023. PMID: 37975835
-
Nearest-neighbor effects on backbone alpha and beta carbon chemical shifts in proteins.J Biomol NMR. 2007 Nov;39(3):247-57. doi: 10.1007/s10858-007-9193-3. J Biomol NMR. 2007. PMID: 17899393
-
Chemical shift-based methods in NMR structure determination.Prog Nucl Magn Reson Spectrosc. 2018 Jun-Aug;106-107:1-25. doi: 10.1016/j.pnmrs.2018.03.002. Epub 2018 Mar 11. Prog Nucl Magn Reson Spectrosc. 2018. PMID: 31047599 Free PMC article. Review.
-
Identification of C-terminal neighbours of amino acid residues without an aliphatic 13Cgamma as an aid to NMR assignments in proteins.J Biomol NMR. 2008 Aug;41(4):191-7. doi: 10.1007/s10858-008-9254-2. Epub 2008 Jul 17. J Biomol NMR. 2008. PMID: 18633715 Review.
Cited by
-
POKY software tools encapsulating assignment strategies for solution and solid-state protein NMR data.J Struct Biol X. 2022 Aug 28;6:100073. doi: 10.1016/j.yjsbx.2022.100073. eCollection 2022. J Struct Biol X. 2022. PMID: 36081577 Free PMC article.
-
NMR Signal Quenching from Bound Biradical Affinity Reagents in DNP Samples.J Phys Chem B. 2017 Dec 7;121(48):10770-10781. doi: 10.1021/acs.jpcb.7b08274. Epub 2017 Nov 29. J Phys Chem B. 2017. PMID: 29116793 Free PMC article.
-
Structural Arrangement within a Peptide Fibril Derived from the Glaucoma-Associated Myocilin Olfactomedin Domain.J Phys Chem B. 2021 Mar 25;125(11):2886-2897. doi: 10.1021/acs.jpcb.0c11460. Epub 2021 Mar 8. J Phys Chem B. 2021. PMID: 33683890 Free PMC article.
-
Resonance assignment of the NMR spectra of disordered proteins using a multi-objective non-dominated sorting genetic algorithm.J Biomol NMR. 2013 Nov;57(3):281-96. doi: 10.1007/s10858-013-9788-9. Epub 2013 Oct 17. J Biomol NMR. 2013. PMID: 24132778 Free PMC article.
-
Molecular conformations and dynamics in the extracellular matrix of mammalian structural tissues: Solid-state NMR spectroscopy approaches.Matrix Biol Plus. 2021 Oct 6;12:100086. doi: 10.1016/j.mbplus.2021.100086. eCollection 2021 Dec. Matrix Biol Plus. 2021. PMID: 34746737 Free PMC article. Review.
References
-
- Cornilescu G, Delaglio F, Bax A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J Biomol NMR. 1999;13:289–302. - PubMed
-
- Creighton TE, editor. Proteins: Structures and molecular properties. W.H. Freeman and Co; New York: 1993.
-
- Eghbalnia HR, Bahrami A, Wang LY, Assadi A, Markley JL. Probabilistic identification of spin systems and their assignments including coil-helix inference as output (PISTACHIO) J Biomol NMR. 2005;32:219–233. - PubMed
-
- Franks WT, Zhou DH, Wylie BJ, Money BG, Graesser DT, Frericks HL, Sahota G, Rienstra CM. Magic-angle spinning solid-state NMR spectroscopy of the beta1 immunoglobulin binding domain of protein G (GB1): 15 N and 13C chemical shift assignments and conformational analysis. J Am Chem Soc. 2005;127:12291–12305. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

