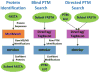
Informatics of protein and posttranslational modification detection via shotgun proteomics
- PMID: 23625403
- PMCID: PMC4012295
- DOI: 10.1007/978-1-62703-360-2_14
Informatics of protein and posttranslational modification detection via shotgun proteomics
Abstract
Frequently, proteomic LC-MS/MS data may contain sets of modifications that evade identification during standard database search. For many laboratories, the standard technique to seek posttranslational modifications (PTMs) adds a short list of specified mass shifts to database search configuration. This technique provides information for only the specified PTMs, takes substantial time to run, and drives false discoveries upward through an exponential expansion of search space. This protocol describes a more structured approach to blind PTM discovery through reducing protein lists, targeting attention to a data-driven list of mass shifts, and seeking the resulting short list of modifications through targeted search.
Figures
References
-
- Eng JK, McCormack AL, Yates JR. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J Am Soc Mass Spectrom. 1994;5:976–989. - PubMed
-
- Yates JR, Eng JK, McCormack AL, Schieltz D. Method to correlate tandem mass spectra of modified peptides to amino acid sequences in the protein database. Anal Chem. 1995;67:1426–1436. - PubMed
-
- Craig R, Beavis RC. A method for reducing the time required to match protein sequences with tandem mass spectra. Rapid Commun Mass Spectrom. 2003;17:2310–2316. - PubMed
-
- Mann M, Wilm M. Error-tolerant identification of peptides in sequence databases by peptide sequence tags. Anal Chem. 1994;66:4390–4399. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous