Engineering A-kinase anchoring protein (AKAP)-selective regulatory subunits of protein kinase A (PKA) through structure-based phage selection
- PMID: 23625929
- PMCID: PMC3682517
- DOI: 10.1074/jbc.M112.447326
Engineering A-kinase anchoring protein (AKAP)-selective regulatory subunits of protein kinase A (PKA) through structure-based phage selection
Abstract
PKA is retained within distinct subcellular environments by the association of its regulatory type II (RII) subunits with A-kinase anchoring proteins (AKAPs). Conventional reagents that universally disrupt PKA anchoring are patterned after a conserved AKAP motif. We introduce a phage selection procedure that exploits high-resolution structural information to engineer RII mutants that are selective for a particular AKAP. Selective RII (RSelect) sequences were obtained for eight AKAPs following competitive selection screening. Biochemical and cell-based experiments validated the efficacy of RSelect proteins for AKAP2 and AKAP18. These engineered proteins represent a new class of reagents that can be used to dissect the contributions of different AKAP-targeted pools of PKA. Molecular modeling and high-throughput sequencing analyses revealed the molecular basis of AKAP-selective interactions and shed new light on native RII-AKAP interactions. We propose that this structure-directed evolution strategy might be generally applicable for the investigation of other protein interaction surfaces.
Keywords: AKAP; Cell Biology; Compartmentalization; Peptide Arrays; Phage Display; Protein Kinase A (PKA); Structure-based Design.
Figures



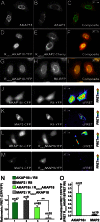


Similar articles
-
AKAP18:PKA-RIIα structure reveals crucial anchor points for recognition of regulatory subunits of PKA.Biochem J. 2016 Jul 1;473(13):1881-94. doi: 10.1042/BCJ20160242. Epub 2016 Apr 21. Biochem J. 2016. PMID: 27102985 Free PMC article.
-
A systematic evaluation of protein kinase A-A-kinase anchoring protein interaction motifs.Biochemistry. 2015 Jan 13;54(1):11-21. doi: 10.1021/bi500721a. Epub 2014 Sep 10. Biochemistry. 2015. PMID: 25097019
-
Single nucleotide polymorphisms alter kinase anchoring and the subcellular targeting of A-kinase anchoring proteins.Proc Natl Acad Sci U S A. 2018 Dec 4;115(49):E11465-E11474. doi: 10.1073/pnas.1816614115. Epub 2018 Nov 19. Proc Natl Acad Sci U S A. 2018. PMID: 30455320 Free PMC article.
-
Anchoring and scaffold proteins for kinases and phosphatases.Recent Prog Horm Res. 1997;52:409-29; discussion 429-30. Recent Prog Horm Res. 1997. PMID: 9238861 Review.
-
Potential therapeutic applications of AKAP disrupting peptides.Clin Sci (Lond). 2020 Dec 23;134(24):3259-3282. doi: 10.1042/CS20201244. Clin Sci (Lond). 2020. PMID: 33346357 Review.
Cited by
-
MaveDB: an open-source platform to distribute and interpret data from multiplexed assays of variant effect.Genome Biol. 2019 Nov 4;20(1):223. doi: 10.1186/s13059-019-1845-6. Genome Biol. 2019. PMID: 31679514 Free PMC article.
-
Molecular Dissection of Neurobeachin Function at Excitatory Synapses.Front Synaptic Neurosci. 2018 Aug 15;10:28. doi: 10.3389/fnsyn.2018.00028. eCollection 2018. Front Synaptic Neurosci. 2018. PMID: 30158865 Free PMC article.
-
Protein kinase A and local signaling in cancer.Biochem J. 2024 Nov 20;481(22):1659-1677. doi: 10.1042/BCJ20230352. Biochem J. 2024. PMID: 39540434 Free PMC article. Review.
-
Signalling scaffolds and local organization of cellular behaviour.Nat Rev Mol Cell Biol. 2015 Apr;16(4):232-44. doi: 10.1038/nrm3966. Epub 2015 Mar 18. Nat Rev Mol Cell Biol. 2015. PMID: 25785716 Free PMC article. Review.
-
Measuring the activity of protein variants on a large scale using deep mutational scanning.Nat Protoc. 2014 Sep;9(9):2267-84. doi: 10.1038/nprot.2014.153. Epub 2014 Aug 28. Nat Protoc. 2014. PMID: 25167058 Free PMC article.
References
-
- Roy J., Cyert M. S. (2009) Cracking the phosphatase code: docking interactions determine substrate specificity. Sci. Signal. 2, re9. - PubMed
-
- Gold M. G., Barford D., Komander D. (2006) Lining the pockets of kinases and phosphatases. Curr. Opin. Struct. Biol. 16, 693–701 - PubMed
-
- Gold M. G., Lygren B., Dokurno P., Hoshi N., McConnachie G., Taskén K., Carlson C. R., Scott J. D., Barford D. (2006) Molecular basis of AKAP specificity for PKA regulatory subunits. Mol. Cell 24, 383–395 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

