HLA engineering of human pluripotent stem cells
- PMID: 23629003
- PMCID: PMC3677304
- DOI: 10.1038/mt.2013.59
HLA engineering of human pluripotent stem cells
Abstract
The clinical use of human pluripotent stem cells and their derivatives is limited by the rejection of transplanted cells due to differences in their human leukocyte antigen (HLA) genes. This has led to the proposed use of histocompatible, patient-specific stem cells; however, the preparation of many different stem cell lines for clinical use is a daunting task. Here, we develop two distinct genetic engineering approaches that address this problem. First, we use a combination of gene targeting and mitotic recombination to derive HLA-homozygous embryonic stem cell (ESC) subclones from an HLA-heterozygous parental line. A small bank of HLA-homozygous stem cells with common haplotypes would match a significant proportion of the population. Second, we derive HLA class I-negative cells by targeted disruption of both alleles of the Beta-2 Microglobulin (B2M) gene in ESCs. Mixed leukocyte reactions and peptide-specific HLA-restricted CD8(+) T cell responses were reduced in class I-negative cells that had undergone differentiation in embryoid bodies. These B2M(-/-) ESCs could act as universal donor cells in applications where the transplanted cells do not express HLA class II genes. Both approaches used adeno-associated virus (AAV) vectors for efficient gene targeting in the absence of potentially genotoxic nucleases, and produced pluripotent, transgene-free cell lines.
Figures

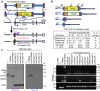


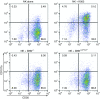
Similar articles
-
Targeted Disruption of the β2-Microglobulin Gene Minimizes the Immunogenicity of Human Embryonic Stem Cells.Stem Cells Transl Med. 2015 Oct;4(10):1234-45. doi: 10.5966/sctm.2015-0049. Epub 2015 Aug 18. Stem Cells Transl Med. 2015. PMID: 26285657 Free PMC article.
-
HLA Class I Depleted hESC as a Source of Hypoimmunogenic Cells for Tissue Engineering Applications.Tissue Eng Part A. 2015 Oct;21(19-20):2559-71. doi: 10.1089/ten.TEA.2015.0105. Epub 2015 Sep 10. Tissue Eng Part A. 2015. PMID: 26218149 Free PMC article.
-
T-Cell Mediated Immune Rejection of Beta-2-Microglobulin Knockout Induced Pluripotent Stem Cell-Derived Kidney Organoids.Stem Cells Transl Med. 2024 Jan 12;13(1):69-82. doi: 10.1093/stcltm/szad069. Stem Cells Transl Med. 2024. PMID: 37843402 Free PMC article.
-
Strategy to Establish Embryo-Derived Pluripotent Stem Cells in Cattle.Int J Mol Sci. 2021 May 9;22(9):5011. doi: 10.3390/ijms22095011. Int J Mol Sci. 2021. PMID: 34065074 Free PMC article. Review.
-
Potential and limitation of HLA-based banking of human pluripotent stem cells for cell therapy.J Immunol Res. 2014;2014:518135. doi: 10.1155/2014/518135. Epub 2014 Jul 9. J Immunol Res. 2014. PMID: 25126584 Free PMC article. Review.
Cited by
-
Regenerative medicine in cardiovascular disease.Regen Ther. 2024 Oct 5;26:859-866. doi: 10.1016/j.reth.2024.09.004. eCollection 2024 Jun. Regen Ther. 2024. PMID: 39430582 Free PMC article. Review.
-
HLA DR Genome Editing with TALENs in Human iPSCs Produced Immune-Tolerant Dendritic Cells.Stem Cells Int. 2021 May 20;2021:8873383. doi: 10.1155/2021/8873383. eCollection 2021. Stem Cells Int. 2021. PMID: 34093711 Free PMC article.
-
Human muscle production in vitro from pluripotent stem cells: Basic and clinical applications.Semin Cell Dev Biol. 2021 Nov;119:39-48. doi: 10.1016/j.semcdb.2021.04.017. Epub 2021 Apr 30. Semin Cell Dev Biol. 2021. PMID: 33941447 Free PMC article. Review.
-
Engineered human pluripotent stem cell-derived natural killer cells: the next frontier for cancer immunotherapy.Blood Sci. 2019 Sep 17;1(1):4-11. doi: 10.1097/BS9.0000000000000023. eCollection 2019 Aug. Blood Sci. 2019. PMID: 35402797 Free PMC article. Review.
-
Strategies for Genetically Engineering Hypoimmunogenic Universal Pluripotent Stem Cells.iScience. 2020 Jun 26;23(6):101162. doi: 10.1016/j.isci.2020.101162. Epub 2020 May 17. iScience. 2020. PMID: 32502965 Free PMC article. Review.
References
-
- Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–872. - PubMed
-
- Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917–1920. - PubMed
-
- Faden RR, Dawson L, Bateman-House AS, Agnew DM, Bok H, Brock DW, et al. Public stem cell banks: considerations of justice in stem cell research and therapy. Hastings Cent Rep. 2003;33:13–27. - PubMed
-
- Taylor CJ, Bolton EM, Pocock S, Sharples LD, Pedersen RA, Bradley JA. Banking on human embryonic stem cells: estimating the number of donor cell lines needed for HLA matching. Lancet. 2005;366:2019–2025. - PubMed
-
- Okita K, Matsumura Y, Sato Y, Okada A, Morizane A, Okamoto S, et al. A more efficient method to generate integration-free human iPS cells. Nat Methods. 2011;8:409–412. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

