Let-7 represses Nr6a1 and a mid-gestation developmental program in adult fibroblasts
- PMID: 23630078
- PMCID: PMC3650230
- DOI: 10.1101/gad.215376.113
Let-7 represses Nr6a1 and a mid-gestation developmental program in adult fibroblasts
Abstract
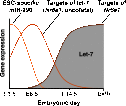
MicroRNAs (miRNAs) are critical to proliferation, differentiation, and development. Here, we characterize gene expression in murine Dicer-null adult mesenchymal stem cell lines, a fibroblast cell type. Loss of Dicer leads to derepression of let-7 targets at levels that exceed 10-fold to 100-fold with increases in transcription. Direct and indirect targets of this miRNA belong to a mid-gestation embryonic program that encompasses known oncofetal genes as well as oncogenes not previously associated with an embryonic state. Surprisingly, this mid-gestation program represents a distinct period that occurs between the pluripotent state of the inner cell mass at embryonic day 3.5 (E3.5) and the induction of let-7 upon differentiation at E10.5. Within this mid-gestation program, we characterize the let-7 target Nr6a1, an embryonic transcriptional repressor that regulates gene expression in adult fibroblasts following miRNA loss. In total, let-7 is required for the continual suppression of embryonic gene expression in adult cells, a mechanism that may underlie its tumor-suppressive function.
Figures






References
-
- Bartel DP, Chen CZ 2004. Micromanagers of gene expression: The potentially widespread influence of metazoan microRNAs. Nat Rev Genet 5: 396–400 - PubMed
-
- Bernstein E, Kim SY, Carmell MA, Murchison EP, Alcorn H, Li MZ, Mills AA, Elledge SJ, Anderson KV, Hannon GJ 2003. Dicer is essential for mouse development. Nat Genet 35: 215–217 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
