Citrus tristeza virus: Evolution of Complex and Varied Genotypic Groups
- PMID: 23630519
- PMCID: PMC3632782
- DOI: 10.3389/fmicb.2013.00093
Citrus tristeza virus: Evolution of Complex and Varied Genotypic Groups
Abstract
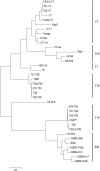
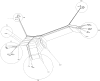
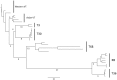
Amongst the Closteroviridae, Citrus tristeza virus (CTV) is almost unique in possessing a number of distinct and characterized strains, isolates of which produce a wide range of phenotype combinations among its different hosts. There is little understanding to connect genotypes to phenotypes, and to complicate matters more, these genotypes are found throughout the world as members of mixed populations within a single host plant. There is essentially no understanding of how combinations of genotypes affect symptom expression and disease severity. We know little about the evolution of the genotypes that have been characterized to date, little about the biological role of their diversity and particularly, about the effects of recombination. Additionally, genotype grouping has not been standardized. In this study we utilized an extensive array of CTV genomic information to classify the major genotypes, and to determine the major evolutionary processes that led to their formation and subsequent retention. Our analyses suggest that three major processes act on these genotypes: (1) ancestral diversification of the major CTV lineages, followed by (2) conservation and co-evolution of the major functional domains within, though not between CTV genotypes, and (3) extensive recombination between lineages that have given rise to new genotypes that have subsequently been retained within the global population. The effects of genotype diversity and host-interaction are discussed, as is a proposal for standardizing the classification of existing and novel CTV genotypes.
Keywords: Citrus tristeza virus; divergence; evolution; genotype; recombination; strain.
Figures

Similar articles
-
Population dynamics of a Florida Citrus tristeza virus isolate and aphid-transmitted subisolates: identification of three genotypic groups and recombinants after aphid transmission.Phytopathology. 2009 Nov;99(11):1297-306. doi: 10.1094/PHYTO-99-11-1297. Phytopathology. 2009. PMID: 19821734
-
Genome analysis of an orange stem pitting citrus tristeza virus isolate reveals a novel recombinant genotype.Virus Res. 2010 Aug;151(2):118-30. doi: 10.1016/j.virusres.2010.03.017. Epub 2010 Apr 8. Virus Res. 2010. PMID: 20381550
-
Analyses of 3' half genome of citrus tristeza virus reveal existence of distinct virus genotypes in citrus growing regions of India.Virusdisease. 2018 Sep;29(3):308-315. doi: 10.1007/s13337-018-0456-2. Epub 2018 Jul 2. Virusdisease. 2018. PMID: 30159365 Free PMC article.
-
Serological and Molecular Detection of Citrus Tristeza Virus: A Review.Microorganisms. 2024 Jul 27;12(8):1539. doi: 10.3390/microorganisms12081539. Microorganisms. 2024. PMID: 39203383 Free PMC article. Review.
-
Walking Together: Cross-Protection, Genome Conservation, and the Replication Machinery of Citrus tristeza virus.Viruses. 2020 Nov 26;12(12):1353. doi: 10.3390/v12121353. Viruses. 2020. PMID: 33256049 Free PMC article. Review.
Cited by
-
Identification of Key Residues Required for RNA Silencing Suppressor Activity of p23 Protein from a Mild Strain of Citrus Tristeza Virus.Viruses. 2019 Aug 25;11(9):782. doi: 10.3390/v11090782. Viruses. 2019. PMID: 31450668 Free PMC article.
-
Minor Variants of Orf1a, p33, and p23 Genes of VT Strain Citrus Tristeza Virus Isolates Show Symptomless Reactions on Sour Orange and Prevent Superinfection of Severe VT Isolates.Viruses. 2023 Sep 30;15(10):2037. doi: 10.3390/v15102037. Viruses. 2023. PMID: 37896814 Free PMC article.
-
A Comprehensive Analysis of Citrus Tristeza Variants of Bhutan and Across the World.Front Microbiol. 2022 Apr 8;13:797463. doi: 10.3389/fmicb.2022.797463. eCollection 2022. Front Microbiol. 2022. PMID: 35464978 Free PMC article.
-
Past and future of a century old Citrus tristeza virus collection: a California citrus germplasm tale.Front Microbiol. 2013 Dec 10;4:366. doi: 10.3389/fmicb.2013.00366. eCollection 2013. Front Microbiol. 2013. PMID: 24339822 Free PMC article.
-
High throughput sequencing from Angolan citrus accessions discloses the presence of emerging CTV strains.Virol J. 2021 Mar 23;18(1):62. doi: 10.1186/s12985-021-01535-x. Virol J. 2021. PMID: 33757535 Free PMC article.
References
-
- Albiach-Marti M. R., Robertson C., Gowda S., Tatineni S., Belliure B., Garnsey S. M., et al. (2010). The pathogenicity determinant of Citrus tristeza virus causing the seedling yellows symptom is located at the 3’-terminal region of the viral genome. Mol. Plant Pathol. 11, 55–6710.1111/j.1364-3703.2009.00572.x - DOI - PMC - PubMed
-
- Ayllon M. A., Navas-Castillo J., Mawassi M., Dawson W. O., Guerri J., Flores R., et al. (1999). New defective RNAs from Citrus tristeza virus: evidence for a replicase-driven template switching mechanism in their generation. J. Gen. Virol. 80, 817–821 - PubMed
-
- Bar-Joseph M., Marcus R., Lee R. F. (1989). The continuing challenge of Citrus tristeza virus control. Annu. Rev. Phytopathol. 27, 291–31610.1146/annurev.py.27.090189.001451 - DOI
-
- Bar-Joseph M., Raccah B., Loebenstein G. (1977). Evaluation of the main variables that affect Citrus tristeza virus transmission by aphids. Proc. Int. Soc. Citriculture 3, 958–961
LinkOut - more resources
Full Text Sources
Other Literature Sources

