Alternative cleavage and polyadenylation: the long and short of it
- PMID: 23632313
- PMCID: PMC3800139
- DOI: 10.1016/j.tibs.2013.03.005
Alternative cleavage and polyadenylation: the long and short of it
Abstract
Cleavage and polyadenylation (C/P) of nascent transcripts is essential for maturation of the 3' ends of most eukaryotic mRNAs. Over the past three decades, biochemical studies have elucidated the machinery responsible for the seemingly simple C/P reaction. Recent genomic analyses have indicated that most eukaryotic genes have multiple cleavage and polyadenylation sites (pAs), leading to transcript isoforms with different coding potentials and/or variable 3' untranslated regions (UTRs). As such, alternative cleavage and polyadenylation (APA) is an important layer of gene regulation impacting mRNA metabolism. Here, we review our current understanding of APA and recent progress in this field.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures

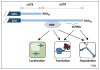

References
-
- Colgan DF, Manley JL. Mechanism and regulation of mRNA polyadenylation. Genes Dev. 1997;11:2755–2766. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

