Origin, evolution and classification of type-3 copper proteins: lineage-specific gene expansions and losses across the Metazoa
- PMID: 23634722
- PMCID: PMC3658974
- DOI: 10.1186/1471-2148-13-96
Origin, evolution and classification of type-3 copper proteins: lineage-specific gene expansions and losses across the Metazoa
Abstract
Background: Tyrosinases, tyrosinase-related proteins, catechol oxidases and hemocyanins comprise the type-3 copper protein family and are involved in a variety of biological processes, including pigment formation, innate immunity and oxygen transport. Although this family is present in the three domains of life, its origin and early evolution are not well understood. Previous analyses of type-3 copper proteins largely have focussed on specific animal and plant phyla.
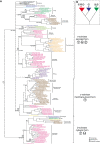
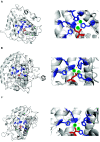
Results: Here, we combine genomic, phylogenetic and structural analyses to show that the original type-3 copper protein possessed a signal peptide and may have been secreted (we designate proteins of this type the α subclass). This ancestral type-3 copper protein gene underwent two duplication events, the first prior to the divergence of the unikont eukaryotic lineages and the second before the diversification of animals. The former duplication gave rise to a cytosolic form (β) and the latter to a membrane-bound form (γ). Structural comparisons reveal that the active site of α and γ forms are covered by aliphatic amino acids, and the β form has a highly conserved aromatic residue in this position. The subsequent evolution of this gene family in modern lineages of multicellular eukaryotes is typified by the loss of one or more of these three subclasses and the lineage-specific expansion of one or both of the remaining subclasses.
Conclusions: The diversity of type-3 copper proteins in animals and other eukaryotes is consistent with two ancient gene duplication events leading to α, β and γ subclasses, followed by the differential loss and expansion of one or more of these subclasses in specific kingdoms and phyla. This has led to many lineage-specific type-3 copper protein repertoires and in some cases the independent evolution of functionally-classified tyrosinases and hemocyanins. For example, the oxygen-carrying hemocyanins in arthropods evolved from a β-subclass tyrosinase, whilst hemocyanins in molluscs and urochordates evolved independently from an α-subclass tyrosinase. Minor conformational changes at the active site of α, β and γ forms can produce type-3 copper proteins with capacities to either carry oxygen (hemocyanins), oxidize diphenols (catechol oxidase) or o-hydroxylate monophenols (tyrosinases) and appear to underlie some functional convergences.
Figures


References
-
- Solomon EI, Baldwin MJ, Lowery MD. Electronic structures of active sites in copper proteins: contributions to reactivity. Chem Rev. 1992;92:521–542. doi: 10.1021/cr00012a003. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

