Pentatricopeptide repeats: modular blocks for building RNA-binding proteins
- PMID: 23635770
- PMCID: PMC3858425
- DOI: 10.4161/rna.24769
Pentatricopeptide repeats: modular blocks for building RNA-binding proteins
Abstract
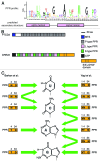
Pentatricopeptide repeat (PPR) proteins control diverse aspects of RNA metabolism across the eukaryotic domain. Recent computational and structural studies have provided new insights into how they recognize RNA, and show that the recognition is sequence-specific and modular. The modular code for RNA-binding by PPR proteins holds great promise for the engineering of new tools to target RNA and identifying RNAs bound by natural PPR proteins.
Keywords: RNA-binding; RNA-protein interactions; pentatricopeptide repeat; repeat proteins; synthetic biology.
Figures

Similar articles
-
An artificial PPR scaffold for programmable RNA recognition.Nat Commun. 2014 Dec 17;5:5729. doi: 10.1038/ncomms6729. Nat Commun. 2014. PMID: 25517350
-
Pentatricopeptide repeats of protein-only RNase P use a distinct mode to recognize conserved bases and structural elements of pre-tRNA.Nucleic Acids Res. 2020 Dec 2;48(21):11815-11826. doi: 10.1093/nar/gkaa627. Nucleic Acids Res. 2020. PMID: 32719843 Free PMC article.
-
RNA-binding specificity landscapes of designer pentatricopeptide repeat proteins elucidate principles of PPR-RNA interactions.Nucleic Acids Res. 2018 Mar 16;46(5):2613-2623. doi: 10.1093/nar/gkx1288. Nucleic Acids Res. 2018. PMID: 29294070 Free PMC article.
-
Human pentatricopeptide proteins: only a few and what do they do?RNA Biol. 2013;10(9):1433-8. doi: 10.4161/rna.24770. Epub 2013 Apr 23. RNA Biol. 2013. PMID: 23635806 Free PMC article. Review.
-
PPR proteins shed a new light on RNase P biology.RNA Biol. 2013;10(9):1457-68. doi: 10.4161/rna.25273. Epub 2013 Jun 19. RNA Biol. 2013. PMID: 23925311 Free PMC article. Review.
Cited by
-
Pentatricopeptide repeat proteins in plants: Cellular functions, action mechanisms, and potential applications.Plant Commun. 2025 Feb 10;6(2):101203. doi: 10.1016/j.xplc.2024.101203. Epub 2024 Dec 5. Plant Commun. 2025. PMID: 39644091 Free PMC article. Review.
-
Mitochondrial ribosomal protein PTCD3 mutations cause oxidative phosphorylation defects with Leigh syndrome.Neurogenetics. 2019 Mar;20(1):9-25. doi: 10.1007/s10048-018-0561-9. Epub 2019 Jan 3. Neurogenetics. 2019. PMID: 30607703
-
Post-Transcriptional Coordination of the Arabidopsis Iron Deficiency Response is Partially Dependent on the E3 Ligases RING DOMAIN LIGASE1 (RGLG1) and RING DOMAIN LIGASE2 (RGLG2).Mol Cell Proteomics. 2015 Oct;14(10):2733-52. doi: 10.1074/mcp.M115.048520. Epub 2015 Aug 7. Mol Cell Proteomics. 2015. PMID: 26253232 Free PMC article.
-
The solution structure of the pentatricopeptide repeat protein PPR10 upon binding atpH RNA.Nucleic Acids Res. 2015 Feb 18;43(3):1918-26. doi: 10.1093/nar/gkv027. Epub 2015 Jan 21. Nucleic Acids Res. 2015. PMID: 25609698 Free PMC article.
-
Biochemical Studies Provide Insights into the Necessity for Multiple Arabidopsis thaliana Protein-Only RNase P Isoenzymes.J Mol Biol. 2019 Feb 1;431(3):615-624. doi: 10.1016/j.jmb.2018.11.004. Epub 2018 Nov 8. J Mol Biol. 2019. PMID: 30414965 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
