Structures of the procapsid and mature virion of enterovirus 71 strain 1095
- PMID: 23637404
- PMCID: PMC3700288
- DOI: 10.1128/JVI.03519-12
Structures of the procapsid and mature virion of enterovirus 71 strain 1095
Abstract
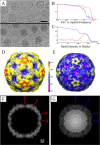
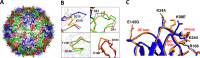
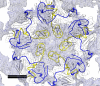
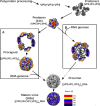
Enterovirus 71 (EV71) is an important emerging human pathogen with a global distribution and presents a disease pattern resembling poliomyelitis with seasonal epidemics that include cases of severe neurological complications, such as acute flaccid paralysis. EV71 is a member of the Picornaviridae family, which consists of icosahedral, nonenveloped, single-stranded RNA viruses. Here we report structures derived from X-ray crystallography and cryoelectron microscopy (cryo-EM) for the 1095 strain of EV71, including a putative precursor in virus assembly, the procapsid, and the mature virus capsid. The cryo-EM map of the procapsid provides new structural information on portions of the capsid proteins VP0 and VP1 that are disordered in the higher-resolution crystal structures. Our structures solved from virus particles in solution are largely in agreement with those from prior X-ray crystallographic studies; however, we observe small but significant structural differences for the 1095 procapsid compared to a structure solved in a previous study (X. Wang, W. Peng, J. Ren, Z. Hu, J. Xu, Z. Lou, X. Li, W. Yin, X. Shen, C. Porta, T. S. Walter, G. Evans, D. Axford, R. Owen, D. J. Rowlands, J. Wang, D. I. Stuart, E. E. Fry, and Z. Rao, Nat. Struct. Mol. Biol. 19:424-429, 2012) for a different strain of EV71. For both EV71 strains, the procapsid is significantly larger in diameter than the mature capsid, unlike in any other picornavirus. Nonetheless, our results demonstrate that picornavirus capsid expansion is possible without RNA encapsidation and that picornavirus assembly may involve an inward radial collapse of the procapsid to yield the native virion.
Figures


References
-
- Schmidt NJ, Lennette EH, Ho HH. 1974. An apparently new enterovirus isolated from patients with disease of the central nervous system. J. Infect. Dis. 129:304–309 - PubMed
-
- Reference deleted.
-
- Solomon T, Lewthwaite P, Perera D, Cardosa MJ, McMinn P, Ooi MH. 2010. Virology, epidemiology, pathogenesis, and control of enterovirus 71. Lancet Infect. Dis. 10:778–790 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

