Visualizing the attack of RNase enzymes on dendriplexes and naked RNA using atomic force microscopy
- PMID: 23637890
- PMCID: PMC3630113
- DOI: 10.1371/journal.pone.0061710
Visualizing the attack of RNase enzymes on dendriplexes and naked RNA using atomic force microscopy
Abstract
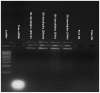
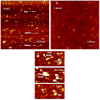
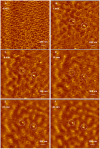
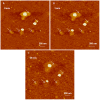
Cationic polymers such as poly(amidoamine), PAMAM, dendrimers have been used to electrostatically complex siRNA molecules forming dendriplexes for enhancing the cytoplasmic delivery of the encapsulated cargo. However, excess PAMAM dendrimers is typically used to protect the loaded siRNA against enzymatic attack, which results in systemic toxicity that hinders the in vivo use of these particles. In this paper, we evaluate the ability of G4 (flexible) and G5 (rigid) dendrimers to complex model siRNA molecules at low +/- ratio of 2/1 upon incubation for 20 minutes and 24 hours. We examine the ability of the formed G4 and G5 dendriplexes to shield the loaded siRNA molecules and protect them from degradation by RNase V1 enzymes using atomic force microscopy (AFM). Results show that G4 and G5 dendrimers form similar hexagonal complexes upon incubation with siRNA molecules for 20 minutes with average full width of 43±19.3 nm and 62±8.3 at half the maximum height, respectively. AFM images show that these G4 and G5 dendriplexes were attacked by RNase V1 enzyme leading to degradation of the exposed RNA molecules that increased with the increase in incubation time. In comparison, incubating G4 and G5 dendrimers with siRNA for 24 hours led to the formation of large particles with average full width of 263±60 nm and 48.3±2.5 nm at half the maximum height, respectively. Both G4 and G5 dendriplexes had a dense central core that proved to shield the loaded RNA molecules from enzymatic attack for up to 60 minutes. These results show the feasibility of formulating G4 and G5 dendriplexes at a low N/P (+/-) ratio that can resist degradation by RNase enzymes, which reduces the risk of inducing non-specific toxicity when used in vivo.
Conflict of interest statement
Figures


References
-
- Hassan A (2006) Potential applications of RNA interference-based therapeutics in the treatment of cardiovascular disease. Recent Pat Cardiovasc Drug Discov 1: 141–149. - PubMed
-
- Koutsilieri E, Rethwilm A, Scheller C (2007) The therapeutic potential of siRNA in gene therapy of neurodegenerative disorders. J Neural Transm Suppl: 43–49. - PubMed
-
- Eccleston A, Eggleston AK (2004) RNA interference. Nature 431: 337–337.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

