Alternative pre-mRNA splicing in neurons: growing up and extending its reach
- PMID: 23648015
- PMCID: PMC3959871
- DOI: 10.1016/j.tig.2013.04.003
Alternative pre-mRNA splicing in neurons: growing up and extending its reach
Abstract
Alternative pre-mRNA splicing determines the protein output of most neuronally expressed genes. Many examples have been described of protein function being modulated by coding changes in different mRNA isoforms. Several recent studies demonstrate that, through the coupling of splicing to other processes of mRNA metabolism, alternative splicing can also act as an on/off switch for gene expression. Other regulated splicing events may determine how an mRNA is utilized in its later cytoplasmic life by changing its localization or translation. These studies make clear that the multiple steps of post-transcriptional gene regulation are strongly linked. Together, these regulatory process play key roles in all aspects of the cell biology of neurons, from their initial differentiation, to their choice of connections, and finally to their function with mature circuits.
Keywords: RNA binding proteins; RNA localization; intron retention; nonsense-mediated mRNA decay; post-transcriptional gene regulation.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures
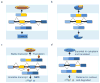
References
-
- Amara SG, et al. Alternative RNA processing in calcitonin gene expression generates mRNAs encoding different polypeptide products. Nature. 1982;298:240–244. - PubMed
-
- Pan Q, et al. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nature genetics. 2008;40:1413–1415. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

