Tracking chromosome evolution in southern African gerbils using flow-sorted chromosome paints
- PMID: 23652816
- PMCID: PMC3721133
- DOI: 10.1159/000350696
Tracking chromosome evolution in southern African gerbils using flow-sorted chromosome paints
Abstract
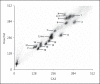
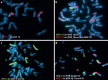
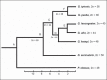
Desmodillus and Gerbilliscus (formerly Tatera) comprise a monophyletic group of gerbils (subfamily Gerbillinae) which last shared an ancestor approximately 8 million years ago; diploid chromosome number variation among the species ranges from 2n = 36 to 2n = 50. In an attempt to shed more light on chromosome evolution and speciation in these rodents, we compared the karyotypes of 7 species, representing 3 genera, based on homology data revealed by chromosome painting with probes derived from flow-sorted chromosomes of the hairy footed gerbil, Gerbillurus paeba (2n = 36). The fluorescent in situ hybridization data revealed remarkable genome conservation: these species share a high proportion of conserved chromosomes, and differences are due to 10 Robertsonian (Rb) rearrangements (3 autapomorphies, 3 synapomorphies and 4 hemiplasies/homoplasies). Our data suggest that chromosome evolution in Desmodillus occurred at a rate of ~1.25 rearrangements per million years (Myr), and that the rate among Gerbilliscus over a time period spanning 8 Myr is also ~1.25 rearrangements/Myr. The recently diverged Gerbillurus (G. tytonis and G. paeba) share an identical karyotype, while Gerbilliscus kempi, G. afra and G. leucogaster differ by 6 Rb rearrangements (a rate of ~1 rearrangement/Myr). Thus, our data suggests a very slow rate of chromosomal evolution in Southern African gerbils.
Copyright © 2013 S. Karger AG, Basel.
Figures



Similar articles
-
Systematics and phylogeny of West African gerbils of the genus Gerbilliscus (Muridae: Gerbillinae) inferred from comparative G- and C-banding chromosomal analyses.Cytogenet Genome Res. 2007;116(4):269-81. doi: 10.1159/000100411. Cytogenet Genome Res. 2007. PMID: 17431325
-
Karyotype evolution in Rhinolophus bats (Rhinolophidae, Chiroptera) illuminated by cross-species chromosome painting and G-banding comparison.Chromosome Res. 2007;15(7):835-48. doi: 10.1007/s10577-007-1167-5. Epub 2007 Oct 1. Chromosome Res. 2007. PMID: 17899409
-
Chromosomal rearrangements and karyotype evolution in carnivores revealed by chromosome painting.Heredity (Edinb). 2012 Jan;108(1):17-27. doi: 10.1038/hdy.2011.107. Epub 2011 Nov 16. Heredity (Edinb). 2012. PMID: 22086079 Free PMC article.
-
Low, complex and probably reticulated chromosome evolution of Sciuromorpha (Rodentia) and Lagomorpha.Cytogenet Genome Res. 2012;137(2-4):218-32. doi: 10.1159/000341379. Epub 2012 Jul 26. Cytogenet Genome Res. 2012. PMID: 22846378 Review.
-
Chromosome evolution in new world monkeys (Platyrrhini).Cytogenet Genome Res. 2012;137(2-4):259-72. doi: 10.1159/000339296. Epub 2012 Jun 12. Cytogenet Genome Res. 2012. PMID: 22699158 Review.
References
-
- Aniskin VM, Benazzou T, Biltueva L, Dobigny G, Granjon L, Volobouev V. Unusually extensive karyotype reorganization in four congeneric Gerbillus species (Muridae: Gerbillinae) Cytogenet Genome Res. 2006;112:131–140. - PubMed
-
- Avise JC, Robinson TJ. Hemiplasy: a new term in the lexicon of phylogenetics. Syst Biol. 2008;57:503–507. - PubMed
-
- Badenhorst D, Dobigny G, Adega F, Chaves R, O'Brien PCM, et al. Chromosomal evolution in Rattini (Muridae, Rodentia) Chromosome Res. 2011;19:709–727. - PubMed
-
- Chevret P, Dobigny G. Systematics and evolution of the subfamily Gerbillinae (Mammalia, Rodentia, Muridae) Mol Phylogenet Evol. 2005;35:674–688. - PubMed
-
- Chimimba CT, Bennett NC. Order Rodentia. In: Skinner JD, Chimimba CT, editors. The Mammals of the Southern African Subregion. Cape Town: Cambridge University Press; 2005. pp. 77–209.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

