Bacterial Diversity Studies Using the 16S rRNA Gene Provide a Powerful Research-Based Curriculum for Molecular Biology Laboratory
- PMID: 23653546
- PMCID: PMC3633119
- DOI: 10.1128/me.3.1.18-25.2002
Bacterial Diversity Studies Using the 16S rRNA Gene Provide a Powerful Research-Based Curriculum for Molecular Biology Laboratory
Abstract
We have developed a ten-week curriculum for molecular biology that uses 16S ribosomal RNA genes to characterize and compare novel bacteria from hot spring communities in Yellowstone National Park. The 16S rRNA approach bypasses selective culture-based methods. Our molecular biology course offered the opportunity for students to learn broadly applicable methods while contributing to a long-term research project. Specifically, students isolated and characterized clones that contained novel 16S rRNA inserts using restriction enzyme, DNA sequencing, and computer-based phylogenetic methods. In both classes, students retrieved novel bacterial 16S rRNA genes, several of which were most similar to Green Nonsulfur bacterial isolates. During class, we evaluated student performance and mastery of skills and concepts using quizzes, formal lab notebooks, and a broad project assignment. For this report, we also assessed student performance alongside data quality and discussed the significance, our goal being to improve both research and teaching methods.
Figures

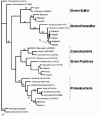

References
-
- Ausubel F, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K, editors. Short protocols in molecular biology. 3rd ed. John Wiley and Sons, Inc; New York, N.Y.: 1997.
-
- Committee on Development of an Addendum to the National Science Education Standards on Scientific Inquiry . Inquiry and the National Science Education Standards, a guide for teaching and learning. National Academy Press; Washington D.C.: 2000.
LinkOut - more resources
Full Text Sources
Miscellaneous

