Gestalt-binding of tropomyosin on actin during thin filament activation
- PMID: 23666668
- PMCID: PMC3773262
- DOI: 10.1007/s10974-013-9342-0
Gestalt-binding of tropomyosin on actin during thin filament activation
Abstract
Our thesis is that thin filament function can only be fully understood and muscle regulation then elucidated if atomic structures of the thin filament are available to reveal the positions of tropomyosin on actin in all physiological states. After all, it is tropomyosin influenced by troponin that regulates myosin-crossbridge cycling on actin and therefore controls contraction in all muscles. In addition, we maintain that a complete appreciation of thin filament activation also requires that the mechanical properties of tropomyosin itself are recognized and then related to the effect of myosin-association on actin. Taking the Gestalt-binding of tropomyosin into account, coupled with our electron microscopy structures and computational chemistry, we propose a comprehensive mechanism for tropomyosin regulatory movement over the actin filament surface that explains the cooperative muscle activation process. In fact, well-known point mutations of critical amino acids on the actin-tropomyosin binding interface disrupt Gestalt-binding and are associated with a number of inherited myopathies. Moreover, dysregulation of tropomyosin may also be a factor that interferes with the gatekeeping operation of non-muscle tropomyosin in the controlling interactions of a wide variety of cellular actin-binding proteins. The clinical relevance of Gestalt-binding is discussed in articles by the Marston and the Gunning groups in this special journal issue devoted to the impact of tropomyosin on biological systems.
Figures

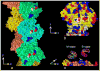


References
-
- Brown JH, Cohen C. Regulation of muscle contraction by tropomyosin and troponin: how structure illuminates function. Adv Protein Chem. 2005;71:121–159. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

