Nucleotide degradation and ribose salvage in yeast
- PMID: 23670538
- PMCID: PMC4039369
- DOI: 10.1038/msb.2013.21
Nucleotide degradation and ribose salvage in yeast
Abstract
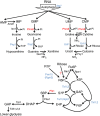
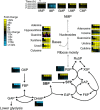
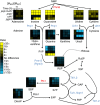
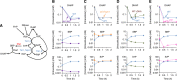
Nucleotide degradation is a universal metabolic capability. Here we combine metabolomics, genetics and biochemistry to characterize the yeast pathway. Nutrient starvation, via PKA, AMPK/SNF1, and TOR, triggers autophagic breakdown of ribosomes into nucleotides. A protein not previously associated with nucleotide degradation, Phm8, converts nucleotide monophosphates into nucleosides. Downstream steps, which involve the purine nucleoside phosphorylase, Pnp1, and pyrimidine nucleoside hydrolase, Urh1, funnel ribose into the nonoxidative pentose phosphate pathway. During carbon starvation, the ribose-derived carbon accumulates as sedoheptulose-7-phosphate, whose consumption by transaldolase is impaired due to depletion of transaldolase's other substrate, glyceraldehyde-3-phosphate. Oxidative stress increases glyceraldehyde-3-phosphate, resulting in rapid consumption of sedoheptulose-7-phosphate to make NADPH for antioxidant defense. Ablation of Phm8 or double deletion of Pnp1 and Urh1 prevent effective nucleotide salvage, resulting in metabolite depletion and impaired survival of starving yeast. Thus, ribose salvage provides means of surviving nutrient starvation and oxidative stress.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures



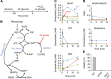
References
-
- Allen J, Davey HM, Broadhurst D, Heald JK, Rowland JJ, Oliver SG, Kell DB (2003) High-throughput classification of yeast mutants for functional genomics using metabolic footprinting. Nat Biotechnol 21: 692–696 - PubMed
-
- Baker CJ, Orlandi EW (1995) Active oxygen in plant pathogenesis. Ann Rev Phytopathol 33: 299–321 - PubMed
-
- Ben-Shem A, de Loubresse NG, Melnikov S, Jenner L, Yusupova G, Yusupov M (2011) The structure of the eukaryotic ribosome at 3.0 angstrom resolution. Science 334: 1524–1529 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

