Molecular evolution of peptidergic signaling systems in bilaterians
- PMID: 23671109
- PMCID: PMC3670399
- DOI: 10.1073/pnas.1219956110
Molecular evolution of peptidergic signaling systems in bilaterians
Abstract
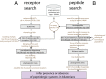
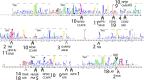
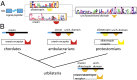
Peptide hormones and their receptors are widespread in metazoans, but the knowledge we have of their evolutionary relationships remains unclear. Recently, accumulating genome sequences from many different species have offered the opportunity to reassess the relationships between protostomian and deuterostomian peptidergic systems (PSs). Here we used sequences of all human rhodopsin and secretin-type G protein-coupled receptors as bait to retrieve potential homologs in the genomes of 15 bilaterian species, including nonchordate deuterostomian and lophotrochozoan species. Our phylogenetic analysis of these receptors revealed 29 well-supported subtrees containing mixed sets of protostomian and deuterostomian sequences. This indicated that many vertebrate and arthropod PSs that were previously thought to be phyla specific are in fact of bilaterian origin. By screening sequence databases for potential peptides, we then reconstructed entire bilaterian peptide families and showed that protostomian and deuterostomian peptides that are ligands of orthologous receptors displayed some similarity at the level of their primary sequence, suggesting an ancient coevolution between peptide and receptor genes. In addition to shedding light on the function of human G protein-coupled receptor PSs, this work presents orthology markers to study ancestral neuron types that were probably present in the last common bilaterian ancestor.
Keywords: GPCR evolution; bilaterian CNS cell types; bilaterian brain evolution; neuropeptide evolution.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Fredriksson R, Lagerström MC, Lundin LG, Schiöth HB. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol Pharmacol. 2003;63(6):1256–1272. - PubMed
-
- Douglass J, Civelli O, Herbert E. Polyprotein gene expression: Generation of diversity of neuroendocrine peptides. Annu Rev Biochem. 1984;53:665–715. - PubMed
-
- Steiner DF. The proprotein convertases. Curr Opin Chem Biol. 1998;2(1):31–39. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

