Direct interaction between the TnsA and TnsB subunits controls the heteromeric Tn7 transposase
- PMID: 23674682
- PMCID: PMC3670325
- DOI: 10.1073/pnas.1305716110
Direct interaction between the TnsA and TnsB subunits controls the heteromeric Tn7 transposase
Abstract
The transposon Tn7 transposase that recognizes the transposon ends and mediates breakage and joining is heteromeric. It contains the Tn7-encoded proteins TnsB, which binds specifically to the transposon ends and carries out breakage and joining at the 3' ends, and TnsA, which carries out breakage at the 5' ends of Tn7. TnsA apparently does not bind specifically to DNA, and we have hypothesized that it is recruited to the ends by interaction with TnsB. In this work, we show that TnsA and TnsB interact directly and identify several TnsA and TnsB amino acids involved in this interaction. We also show that TnsA can stimulate two key activities of TnsB, specific binding to the ends and pairing of the Tn7 ends. The ends of Tn7 are structurally asymmetric (i.e., contain different numbers of TnsB-binding sites), and Tn7 also is functionally asymmetric, inserting into its specific target site, attachment site attTn7 (attTn7) in a single orientation. Moreover, Tn7 elements containing two Tn7 right ends can transpose, but elements with two Tn7 left ends cannot. We show here that TnsA + TnsB are unable to pair the ends of a Tn7 element containing two Tn7 left ends. This pairing defect likely contributes to the inability of Tn7 elements with two Tn7 left ends to transpose.
Keywords: asymmetric transposon ends; protein-DNA interaction; protein–protein interaction; transposition.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

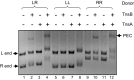
References
-
- Craig NL, Craigie R, Gellert M, Lambowitz A. Mobile DNA II. Washington, DC: ASM; 2002.
-
- Peters JE, Craig NL. Tn7: Smarter than we thought. Nat Rev Mol Cell Biol. 2001;2(11):806–814. - PubMed
-
- Li Z, Craig NL, Peters JE. 2013. in Bacterial Integrative Mobile Genetic Elements, eds Roberts A, Mullany P (Landes Bioscience, London), pp. 1–32.
-
- May EW, Craig NL. Switching from cut-and-paste to replicative Tn7 transposition. Science. 1996;272(5260):401–404. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

