A unique spumavirus Gag N-terminal domain with functional properties of orthoretroviral matrix and capsid
- PMID: 23675305
- PMCID: PMC3649970
- DOI: 10.1371/journal.ppat.1003376
A unique spumavirus Gag N-terminal domain with functional properties of orthoretroviral matrix and capsid
Abstract
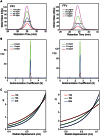
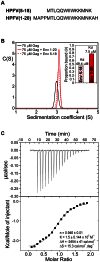
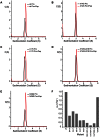
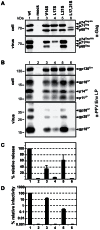
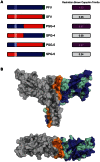
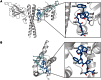
The Spumaretrovirinae, or foamyviruses (FVs) are complex retroviruses that infect many species of monkey and ape. Although FV infection is apparently benign, trans-species zoonosis is commonplace and has resulted in the isolation of the Prototypic Foamy Virus (PFV) from human sources and the potential for germ-line transmission. Despite little sequence homology, FV and orthoretroviral Gag proteins perform equivalent functions, including genome packaging, virion assembly, trafficking and membrane targeting. In addition, PFV Gag interacts with the FV Envelope (Env) protein to facilitate budding of infectious particles. Presently, there is a paucity of structural information with regards FVs and it is unclear how disparate FV and orthoretroviral Gag molecules share the same function. Therefore, in order to probe the functional overlap of FV and orthoretroviral Gag and learn more about FV egress and replication we have undertaken a structural, biophysical and virological study of PFV-Gag. We present the crystal structure of a dimeric amino terminal domain from PFV, Gag-NtD, both free and in complex with the leader peptide of PFV Env. The structure comprises a head domain together with a coiled coil that forms the dimer interface and despite the shared function it is entirely unrelated to either the capsid or matrix of Gag from other retroviruses. Furthermore, we present structural, biochemical and virological data that reveal the molecular details of the essential Gag-Env interaction and in addition we also examine the specificity of Trim5α restriction of PFV. These data provide the first information with regards to FV structural proteins and suggest a model for convergent evolution of gag genes where structurally unrelated molecules have become functionally equivalent.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Structure of a Spumaretrovirus Gag Central Domain Reveals an Ancient Retroviral Capsid.PLoS Pathog. 2016 Nov 9;12(11):e1005981. doi: 10.1371/journal.ppat.1005981. eCollection 2016 Nov. PLoS Pathog. 2016. PMID: 27829070 Free PMC article.
-
An N-terminal domain helical motif of Prototype Foamy Virus Gag with dual functions essential for particle egress and viral infectivity.Retrovirology. 2013 Apr 25;10:45. doi: 10.1186/1742-4690-10-45. Retrovirology. 2013. PMID: 23618494 Free PMC article.
-
Mutagenesis of N-terminal residues of feline foamy virus Gag reveals entirely distinct functions during capsid formation, particle assembly, Gag processing and budding.Retrovirology. 2016 Aug 22;13(1):57. doi: 10.1186/s12977-016-0291-8. Retrovirology. 2016. PMID: 27549192 Free PMC article.
-
The foamy virus Gag proteins: what makes them different?Viruses. 2013 Mar 26;5(4):1023-41. doi: 10.3390/v5041023. Viruses. 2013. PMID: 23531622 Free PMC article. Review.
-
The Unique, the Known, and the Unknown of Spumaretrovirus Assembly.Viruses. 2021 Jan 13;13(1):105. doi: 10.3390/v13010105. Viruses. 2021. PMID: 33451128 Free PMC article. Review.
Cited by
-
Foamy Viruses, Bet, and APOBEC3 Restriction.Viruses. 2021 Mar 18;13(3):504. doi: 10.3390/v13030504. Viruses. 2021. PMID: 33803830 Free PMC article. Review.
-
Identification of Cartilaginous Fish Endogenous Foamy Virus Rooting to Vertebrate Counterparts.J Virol. 2023 Feb 28;97(2):e0181622. doi: 10.1128/jvi.01816-22. Epub 2023 Jan 18. J Virol. 2023. PMID: 36651746 Free PMC article.
-
Novel Host Protein TBC1D16, a GTPase Activating Protein of Rab5C, Inhibits Prototype Foamy Virus Replication.Front Immunol. 2021 Jul 22;12:658660. doi: 10.3389/fimmu.2021.658660. eCollection 2021. Front Immunol. 2021. PMID: 34367131 Free PMC article.
-
Distinct Particle Morphologies Revealed through Comparative Parallel Analyses of Retrovirus-Like Particles.J Virol. 2016 Aug 26;90(18):8074-84. doi: 10.1128/JVI.00666-16. Print 2016 Sep 15. J Virol. 2016. PMID: 27356903 Free PMC article.
-
Structural and Functional Aspects of Foamy Virus Protease-Reverse Transcriptase.Viruses. 2019 Jul 2;11(7):598. doi: 10.3390/v11070598. Viruses. 2019. PMID: 31269675 Free PMC article. Review.
References
-
- Lecellier CH, Saib A (2000) Foamy viruses: between retroviruses and pararetroviruses. Virology 271: 1–8. - PubMed
-
- Herchenroder O, Renne R, Loncar D, Cobb EK, Murthy KK, et al. (1994) Isolation, cloning, and sequencing of simian foamy viruses from chimpanzees (SFVcpz): high homology to human foamy virus (HFV). Virology 201: 187–199. - PubMed
-
- Bieniasz PD, Rethwilm A, Pitman R, Daniel MD, Chrystie I, et al. (1995) A comparative study of higher primate foamy viruses, including a new virus from a gorilla. Virology 207: 217–228. - PubMed
-
- Broussard SR, Comuzzie AG, Leighton KL, Leland MM, Whitehead EM, et al. (1997) Characterization of new simian foamy viruses from African nonhuman primates. Virology 237: 349–359. - PubMed
-
- Saib A (2003) Non-primate foamy viruses. Curr Top Microbiol Immunol 277: 197–211. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

