Molecular determinants of regulatory T cell development: the essential roles of epigenetic changes
- PMID: 23675373
- PMCID: PMC3650462
- DOI: 10.3389/fimmu.2013.00106
Molecular determinants of regulatory T cell development: the essential roles of epigenetic changes
Abstract
Regulatory T (Treg) cells constitute a distinct T cell subset, which plays a key role in immune tolerance and homeostasis. The transcription factor Foxp3 controls a substantial part of Treg cell development and function. Yet its expression alone is insufficient for conferring developmental and functional characteristics of Treg cells. There is accumulating evidence that concurrent induction of Treg-specific epigenetic changes and Foxp3 expression is crucial for lineage specification and functional stability of Treg cells. This review discusses recent progress in our understanding of molecular features of Treg cells, in particular, the molecular basis of how a population of developing T cells is driven to the Treg cell lineage and how its function is stably maintained.
Keywords: DNA methylation; Foxp3; adaptability; epigenetics; lineage specification; plasticity; regulatory T cells.
Figures

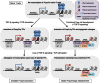
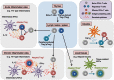
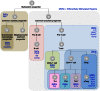
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

