Dendritic GluN2A synthesis mediates activity-induced NMDA receptor insertion
- PMID: 23678131
- PMCID: PMC3684268
- DOI: 10.1523/JNEUROSCI.0289-13.2013
Dendritic GluN2A synthesis mediates activity-induced NMDA receptor insertion
Abstract
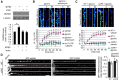
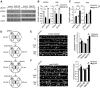
Long-term synaptic plasticity involves changes in the expression and membrane insertion of cell-surface proteins. Interestingly, the mRNAs encoding many cell-surface proteins are localized to dendrites, but whether dendritic protein synthesis is required for activity-induced surface expression of specific proteins is unknown. Herein, we used microfluidic devices to demonstrate that dendritic protein synthesis is necessary for activity-induced insertion of GluN2A-containing NMDA receptors in rat hippocampal neurons. Furthermore, visualization of activity-induced local translation of GluN2A mRNA and membrane insertion of GluN2A protein in dendrites was directly observed and shown to depend on a 3' untranslated region cytoplasmic polyadenylation element and its associated translation complex. These findings uncover a novel mechanism for cytoplasmic polyadenylation element-mediated posttranscriptional regulation of GluN2A mRNA to control NMDA receptor surface expression during synaptic plasticity.
Figures





References
-
- Anji A, Kumari M. A cis-acting region in the N-methyl-d-aspartate R1 3′-untranslated region interacts with the novel RNA-binding proteins beta subunit of alpha glucosidase II and annexin A2: effect of chronic ethanol exposure in vivo. Eur J Neurosci. 2011;34:1200–1211. doi: 10.1111/j.1460-9568.2011.07857.x. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
