ER-associated mitochondrial division links the distribution of mitochondria and mitochondrial DNA in yeast
- PMID: 23682313
- PMCID: PMC3654481
- DOI: 10.7554/eLife.00422
ER-associated mitochondrial division links the distribution of mitochondria and mitochondrial DNA in yeast
Abstract
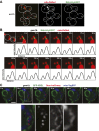
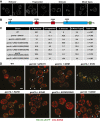
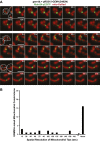
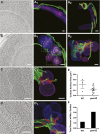
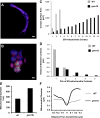
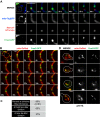
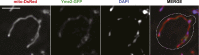
Mitochondrial division is important for mitochondrial distribution and function. Recent data have demonstrated that ER-mitochondria contacts mark mitochondrial division sites, but the molecular basis and functions of these contacts are not understood. Here we show that in yeast, the ER-mitochondria tethering complex, ERMES, and the highly conserved Miro GTPase, Gem1, are spatially and functionally linked to ER-associated mitochondrial division. Gem1 acts as a negative regulator of ER-mitochondria contacts, an activity required for the spatial resolution and distribution of newly generated mitochondrial tips following division. Previous data have demonstrated that ERMES localizes with a subset of actively replicating mitochondrial nucleoids. We show that mitochondrial division is spatially linked to nucleoids and that a majority of these nucleoids segregate prior to division, resulting in their distribution into newly generated tips in the mitochondrial network. Thus, we postulate that ER-associated division serves to link the distribution of mitochondria and mitochondrial nucleoids in cells. DOI:http://dx.doi.org/10.7554/eLife.00422.001.
Keywords: ERMES; Gem1; Miro; S. cerevisiae; mitochondria; mitochondrial DNA.
Conflict of interest statement
JN, Reviewing editor,
The other authors declare that no competing interests exist.
Figures





Comment in
-
Make or break for mitochondria.Elife. 2013 May 14;2:e00804. doi: 10.7554/eLife.00804. Elife. 2013. PMID: 23682317 Free PMC article.
References
-
- Brachmann CB, Davies A, Cost GJ, Caputo E, LI J, Hieter P, et al. 1998. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14:115–32 doi: 10.1002/(SICI)1097-0061(19980130)14:2<115::AID-YEA204>3.0.CO;2-2 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

