L2, the minor capsid protein of papillomavirus
- PMID: 23689062
- PMCID: PMC3770800
- DOI: 10.1016/j.virol.2013.04.017
L2, the minor capsid protein of papillomavirus
Abstract
The capsid protein L2 plays major roles in both papillomavirus assembly and the infectious process. While L1 forms the majority of the capsid and can self-assemble into empty virus-like particles (VLPs), L2 is a minor capsid component and lacks the capacity to form VLPs. However, L2 co-assembles with L1 into VLPs, enhancing their assembly. L2 also facilitates encapsidation of the ∼8 kbp circular and nucleosome-bound viral genome during assembly of the non-enveloped T=7d virions in the nucleus of terminally differentiated epithelial cells, although, like L1, L2 is not detectably expressed in infected basal cells. With respect to infection, L2 is not required for particles to bind to and enter cells. However L2 must be cleaved by furin for endosome escape. L2 then travels with the viral genome to the nucleus, wherein it accumulates at ND-10 domains. Here, we provide an overview of the biology of L2.
Keywords: Daxx; E2; Encapsidation; Furin; HPV; Infection; L1; L2; Minor capsid protein; ND-10; Papillomavirus; SP100.
© 2013 Elsevier Inc. All rights reserved.
Conflict of interest statement
Figures



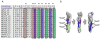

References
-
- Becker KA, Florin L, Sapp C, Sapp M. Dissection of human papillomavirus type 33 L2 domains involved in nuclear domains (ND) 10 homing and reorganization. Virology. 2003;314:161–167. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

