Thermostable group II intron reverse transcriptase fusion proteins and their use in cDNA synthesis and next-generation RNA sequencing
- PMID: 23697550
- PMCID: PMC3683930
- DOI: 10.1261/rna.039743.113
Thermostable group II intron reverse transcriptase fusion proteins and their use in cDNA synthesis and next-generation RNA sequencing
Abstract
Mobile group II introns encode reverse transcriptases (RTs) that function in intron mobility ("retrohoming") by a process that requires reverse transcription of a highly structured, 2-2.5-kb intron RNA with high processivity and fidelity. Although the latter properties are potentially useful for applications in cDNA synthesis and next-generation RNA sequencing (RNA-seq), group II intron RTs have been difficult to purify free of the intron RNA, and their utility as research tools has not been investigated systematically. Here, we developed general methods for the high-level expression and purification of group II intron-encoded RTs as fusion proteins with a rigidly linked, noncleavable solubility tag, and we applied them to group II intron RTs from bacterial thermophiles. We thus obtained thermostable group II intron RT fusion proteins that have higher processivity, fidelity, and thermostability than retroviral RTs, synthesize cDNAs at temperatures up to 81°C, and have significant advantages for qRT-PCR, capillary electrophoresis for RNA-structure mapping, and next-generation RNA sequencing. Further, we find that group II intron RTs differ from the retroviral enzymes in template switching with minimal base-pairing to the 3' ends of new RNA templates, making it possible to efficiently and seamlessly link adaptors containing PCR-primer binding sites to cDNA ends without an RNA ligase step. This novel template-switching activity enables facile and less biased cloning of nonpolyadenylated RNAs, such as miRNAs or protein-bound RNA fragments. Our findings demonstrate novel biochemical activities and inherent advantages of group II intron RTs for research, biotechnological, and diagnostic methods, with potentially wide applications.
Keywords: miRNA; next-generation sequencing; qRT-PCR; retrovirus; transcriptome.
Figures




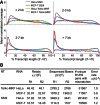



References
-
- Arezi B, Hogrefe HH 2007. Escherichia coli DNA polymerase III ε subunit increases Moloney murine leukemia virus reverse transcriptase fidelity and accuracy of RT–PCR procedures. Anal Biochem 360: 84–91 - PubMed
-
- Baranauskas A, Paliksa S, Alzbutas G, Vaitkevicius M, Lubiene J, Letukiene V, Burinskas S, Sasnauskas G, Skirgaila R 2012. Generation and characterization of new highly thermostable and processive M-MuLV reverse transcriptase variants. Protein Eng Des Sel 25: 657–668 - PubMed
-
- Beckman RA, Mildvan AS, Loeb LA 1985. On the fidelity of DNA replication: Manganese mutagenesis in vitro. Biochemistry 24: 5810–5817 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
