MicroRNA-185 oscillation controls circadian amplitude of mouse Cryptochrome 1 via translational regulation
- PMID: 23699394
- PMCID: PMC3708730
- DOI: 10.1091/mbc.E12-12-0849
MicroRNA-185 oscillation controls circadian amplitude of mouse Cryptochrome 1 via translational regulation
Abstract
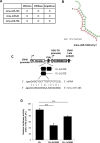
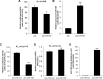
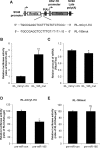
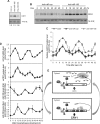
Mammalian circadian rhythm is observed not only at the suprachiasmatic nucleus, a master pacemaker, but also throughout the peripheral tissues. Investigation of the regulation of clock gene expression has mainly focused on transcriptional and posttranslational modifications, and little is known about the posttranscriptional regulation of these genes. In the present study, we investigate the role of microRNAs (miRNAs) in the posttranscriptional regulation of the 3'-untranslated region (UTR) of the mouse Cryptochrome 1 (mCry1) gene. Knockdown of Drosha, Dicer, or Argonaute2 increased mCry1-3'UTR reporter activity. The presence of the miRNA recognition element of mCry1 that is important for miR-185 binding decreased mCRY1 protein, but not mRNA, level. Cytoplasmic miR-185 levels were nearly antiphase to mCRY1 protein levels. Furthermore, miR-185 knockdown elevated the amplitude of mCRY1 protein oscillation. Our results suggest that miR-185 plays a role in the fine-tuned regulation of mCRY1 protein expression by controlling the rhythmicity of mCry1 mRNA translation.
Figures

Similar articles
-
Circadian amplitude of cryptochrome 1 is modulated by mRNA stability regulation via cytoplasmic hnRNP D oscillation.Mol Cell Biol. 2010 Jan;30(1):197-205. doi: 10.1128/MCB.01154-09. Mol Cell Biol. 2010. PMID: 19858287 Free PMC article.
-
microRNA-25 as a novel modulator of circadian Period2 gene oscillation.Exp Mol Med. 2020 Sep;52(9):1614-1626. doi: 10.1038/s12276-020-00496-5. Epub 2020 Sep 23. Exp Mol Med. 2020. PMID: 32968200 Free PMC article.
-
AUF1 contributes to Cryptochrome1 mRNA degradation and rhythmic translation.Nucleic Acids Res. 2014 Apr;42(6):3590-606. doi: 10.1093/nar/gkt1379. Epub 2014 Jan 13. Nucleic Acids Res. 2014. PMID: 24423872 Free PMC article.
-
Regulation of microRNA biogenesis and function.Thromb Haemost. 2012 Apr;107(4):605-10. doi: 10.1160/TH11-12-0836. Epub 2012 Feb 8. Thromb Haemost. 2012. PMID: 22318703 Review.
-
Regulation of mammalian transcription and splicing by Nuclear RNAi.Nucleic Acids Res. 2016 Jan 29;44(2):524-37. doi: 10.1093/nar/gkv1305. Epub 2015 Nov 26. Nucleic Acids Res. 2016. PMID: 26612865 Free PMC article. Review.
Cited by
-
Systems Biology-Derived Discoveries of Intrinsic Clocks.Front Neurol. 2017 Feb 6;8:25. doi: 10.3389/fneur.2017.00025. eCollection 2017. Front Neurol. 2017. PMID: 28220104 Free PMC article. Review.
-
Nutrigenetics and Nutrimiromics of the Circadian System: The Time for Human Health.Int J Mol Sci. 2016 Feb 26;17(3):299. doi: 10.3390/ijms17030299. Int J Mol Sci. 2016. PMID: 26927084 Free PMC article. Review.
-
MicroRNA: A Key Player for the Interplay of Circadian Rhythm Abnormalities, Sleep Disorders and Neurodegenerative Diseases.Clocks Sleep. 2020 Jul 23;2(3):282-307. doi: 10.3390/clockssleep2030022. eCollection 2020 Sep. Clocks Sleep. 2020. PMID: 33089205 Free PMC article. Review.
-
MicroRNAs shape circadian hepatic gene expression on a transcriptome-wide scale.Elife. 2014 May 27;3:e02510. doi: 10.7554/eLife.02510. Elife. 2014. PMID: 24867642 Free PMC article.
-
Out-of-Equilibrium ceRNA Crosstalk.Methods Mol Biol. 2025;2883:167-193. doi: 10.1007/978-1-0716-4290-0_8. Methods Mol Biol. 2025. PMID: 39702709
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

