Cellular metabolic and autophagic pathways: traffic control by redox signaling
- PMID: 23702245
- PMCID: PMC3729625
- DOI: 10.1016/j.freeradbiomed.2013.05.014
Cellular metabolic and autophagic pathways: traffic control by redox signaling
Abstract
It has been established that the key metabolic pathways of glycolysis and oxidative phosphorylation are intimately related to redox biology through control of cell signaling. Under physiological conditions glucose metabolism is linked to control of the NADH/NAD redox couple, as well as providing the major reductant, NADPH, for thiol-dependent antioxidant defenses. Retrograde signaling from the mitochondrion to the nucleus or cytosol controls cell growth and differentiation. Under pathological conditions mitochondria are targets for reactive oxygen and nitrogen species and are critical in controlling apoptotic cell death. At the interface of these metabolic pathways, the autophagy-lysosomal pathway functions to maintain mitochondrial quality and generally serves an important cytoprotective function. In this review we will discuss the autophagic response to reactive oxygen and nitrogen species that are generated from perturbations of cellular glucose metabolism and bioenergetic function.
Keywords: 4-hydroxy-2-nonenal; AMP kinase; AMPK; Aging; Ambra; Atg; Autophagy; B-cell lymphoma 2; BH3; BNIP; Bcl-2; Bcl-2 homology domain 3; Bcl-2/adenovirus E18 19-kDa-interacting protein; CMA; Cardiovascular disease; Cellular bioenergetics; Diabetes; ER; ERK; ESCRT3; Free radicals; G6PD; GAPDH; GSH; GSSG; Glucose; Glutathione; Glycolysis; HIF; HNE; Hsc70; IRE; JNK; LC3; MEF; Mitochondria; Mitophagy; NADPH oxidase; NIP3-like X; NIX; NOX; Neurodegeneration; Oxidative stress; PI3K; PI3P; PINK1; PPP; PTEN-induced kinase 1; RNS; ROS; Rab5; Rac1; Ras-associated protein; Ras-related C3 botulinum toxin substrate 1; Redox signaling; SNARE; TIGAR; TP53-induced glycolysis and apoptosis regulator; TSC; ULK1 and ULK2; UV radiation resistance-associated gene protein; UVRAG; VDAC; Vacuolar protein sorting.; Vps; activating molecule in beclin 1-regulated autophagy; autophagy-related gene; c-Jun N-terminal kinase; chaperone-mediated autophagy; eNOS; endoplasmic reticulum; endosomal sorting complexes required for transport 3; endothelial nitric oxide synthase; extracellular signal-regulated kinase; glucose-6-phosphate dehydrogenase; glutathione; glyceraldehyde-3-phosphate dehydrogenase; heat shock cognate 70; hypoxia-inducible factor; inositol requiring enzyme; mTOR; mammalian target of rapamycin; microtubule-associated protein 1 light-chain subunit 3; mouse embryonic fibroblast; oxidized glutathione; pentose phosphate pathway; phosphatidylinositol 3-phosphate; phosphoinositide 3-kinase; reactive nitrogen species; reactive oxygen species; soluble N-ethylmaleimide-sensitive factor attachment protein receptor; tuberous sclerosis complex; uncoordinated family member-51-like kinases 1 and 2; voltage-dependent anion channel.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures

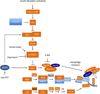
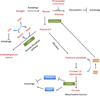


References
-
- Pan T, Kondo S, Le W, Jankovic J. The role of autophagy-lysosome pathway in neurodegeneration associated with Parkinson's disease. Brain. 2008;131:1969–1978. - PubMed
-
- Trushina E, McMurray CT. Oxidative stress and mitochondrial dysfunction in neurodegenerative diseases. Neuroscience. 2007;145:1233–1248. - PubMed
-
- Beal MF. Mitochondria take center stage in aging and neurodegeneration. Ann Neurol. 2005;58:495–505. - PubMed
-
- Levine B. Cell biology: autophagy and cancer. Nature. 2007;446:745–747. - PubMed
-
- Nishida K, Kyoi S, Yamaguchi O, Sadoshima J, Otsu K. The role of autophagy in the heart. Cell Death Differ. 2009;16:31–38. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

