A Raman-based endoscopic strategy for multiplexed molecular imaging
- PMID: 23703909
- PMCID: PMC3690865
- DOI: 10.1073/pnas.1211309110
A Raman-based endoscopic strategy for multiplexed molecular imaging
Abstract
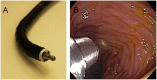
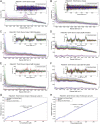
Endoscopic imaging is an invaluable diagnostic tool allowing minimally invasive access to tissues deep within the body. It has played a key role in screening colon cancer and is credited with preventing deaths through the detection and removal of precancerous polyps. However, conventional white-light endoscopy offers physicians structural information without the biochemical information that would be advantageous for early detection and is essential for molecular typing. To address this unmet need, we have developed a unique accessory, noncontact, fiber optic-based Raman spectroscopy device that has the potential to provide real-time, multiplexed functional information during routine endoscopy. This device is ideally suited for detection of functionalized surface-enhanced Raman scattering (SERS) nanoparticles as molecular imaging contrast agents. This device was designed for insertion through a clinical endoscope and has the potential to detect and quantify the presence of a multiplexed panel of tumor-targeting SERS nanoparticles. Characterization of the Raman instrument was performed with SERS particles on excised human tissue samples, and it has shown unsurpassed sensitivity and multiplexing capabilities, detecting 326-fM concentrations of SERS nanoparticles and unmixing 10 variations of colocalized SERS nanoparticles. Another unique feature of our noncontact Raman endoscope is that it has been designed for efficient use over a wide range of working distances from 1 to 10 mm. This is necessary to accommodate for imperfect centering during endoscopy and the nonuniform surface topology of human tissue. Using this endoscope as a key part of a multiplexed detection approach could allow endoscopists to distinguish between normal and precancerous tissues rapidly and to identify flat lesions that are otherwise missed.
Keywords: colonoscopy; optical.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
-
- Raman CV. A new radiation. Indian J Phys. 1928;2:387–398.
-
- Crow P, et al. Assessment of fiberoptic near-infrared raman spectroscopy for diagnosis of bladder and prostate cancer. Urology. 2005;65(6):1126–1130. - PubMed
-
- Kendall C, et al. Vibrational spectroscopy: A clinical tool for cancer diagnostics. Analyst (Lond) 2009;134(6):1029–1045. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

