Reversible, long-range radical transfer in E. coli class Ia ribonucleotide reductase
- PMID: 23730940
- PMCID: PMC3823682
- DOI: 10.1021/ar4000407
Reversible, long-range radical transfer in E. coli class Ia ribonucleotide reductase
Abstract
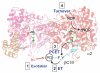
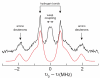
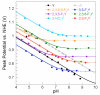
Ribonucleotide reductases (RNRs) catalyze the conversionof nucleotides to 2'-deoxynucleotides and are classified on the basis of the metallo-cofactor used to conduct this chemistry. The class Ia RNRs initiate nucleotide reduction when a stable diferric-tyrosyl radical (Y•, t1/2 of 4 days at 4 °C) cofactor in the β2 subunit transiently oxidizes a cysteine to a thiyl radical (S•) in the active site of the α2 subunit. In the active α2β2 complex of the class Ia RNR from E. coli , researchers have proposed that radical hopping occurs reversibly over 35 Å along a specific pathway comprised of redox-active aromatic amino acids: Y122• ↔ [W48?] ↔ Y356 in β2 to Y731 ↔ Y730 ↔ C439 in α2. Each step necessitates a proton-coupled electron transfer (PCET). Protein conformational changes constitute the rate-limiting step in the overall catalytic scheme and kinetically mask the detailed chemistry of the PCET steps. Technology has evolved to allow the site-selective replacement of the four pathway tyrosines with unnatural tyrosine analogues. Rapid kinetic techniques combined with multifrequency electron paramagnetic resonance, pulsed electron-electron double resonance, and electron nuclear double resonance spectroscopies have facilitated the analysis of stable and transient radical intermediates in these mutants. These studies are beginning to reveal the mechanistic underpinnings of the radical transfer (RT) process. This Account summarizes recent mechanistic studies on mutant E. coli RNRs containing the following tyrosine analogues: 3,4-dihydroxyphenylalanine (DOPA) or 3-aminotyrosine (NH2Y), both thermodynamic radical traps; 3-nitrotyrosine (NO2Y), a thermodynamic barrier and probe of local environmental perturbations to the phenolic pKa; and fluorotyrosines (FnYs, n = 2 or 3), dual reporters on local pKas and reduction potentials. These studies have established the existence of a specific pathway spanning 35 Å within a globular α2β2 complex that involves one stable (position 122) and three transient (positions 356, 730, and 731) Y•s. Our results also support that RT occurs by an orthogonal PCET mechanism within β2, with Y122• reduction accompanied by proton transfer from an Fe1-bound water in the diferric cluster and Y356 oxidation coupled to an off-pathway proton transfer likely involving E350. In α2, RT likely occurs by a co-linear PCET mechanism, based on studies of light-initiated radical propagation from photopeptides that mimic the β2 subunit to the intact α2 subunit and on [(2)H]-ENDOR spectroscopic analysis of the hydrogen-bonding environment surrounding a stabilized NH2Y• formed at position 730. Additionally, studies on the thermodynamics of the RT pathway reveal that the relative reduction potentials decrease according to Y122 < Y356 < Y731 ≈ Y730 ≤ C439, and that the pathway in the forward direction is thermodynamically unfavorable. C439 oxidation is likely driven by rapid, irreversible loss of water during the nucleotide reduction process. Kinetic studies of radical intermediates reveal that RT is gated by conformational changes that occur on the order of >100 s(-1) in addition to the changes that are rate-limiting in the wild-type enzyme (∼10 s(-1)). The rate constant of one of the PCET steps is ∼10(5) s(-1), as measured in photoinitiated experiments.
Figures









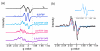

References
-
- Stubbe J, van der Donk WA. Protein radicals in enzyme catalysis. Chem. Rev. 1998;98:705–762. and references therein. - PubMed
-
- Uhlin U, Eklund H. Structure of ribonucleotide reductase protein R1. Nature. 1994;370:533–539. - PubMed
-
- Licht S, Stubbe J. Mechanistic investigations of ribonucleotide reductases. Compr. Nat. Prod. Chem. 1999;5:163–203. and references therein.
-
- Climent I, Sjöberg BM, Huang CY. Carboxyl-terminal peptides as probes for Escherichia coli ribonucleotide reductase subunit interaction: kinetic analysis of inhibition studies. Biochemistry. 1991;30:5164–5171. - PubMed
-
- Ge J, Yu G, Ator MA, Stubbe J. Pre-steady-state and steady-state kinetic analysis of E. coli class I ribonucleotide reductase. Biochemistry. 2003;42:10071–10083. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

