p28, a first in class peptide inhibitor of cop1 binding to p53
- PMID: 23736031
- PMCID: PMC3694247
- DOI: 10.1038/bjc.2013.266
p28, a first in class peptide inhibitor of cop1 binding to p53
Abstract
Background: A 28 amino-acid (aa) cell-penetrating peptide (p28) derived from azurin, a redox protein secreted from the opportunistic pathogen Pseudomonas aeruginosa, produces a post-translational increase in p53 in cancer cells by inhibiting its ubiquitination.
Methods: In silico computational simulations were used to predict motifs within the p53 DNA-binding domain (DBD) as potential sites for p28 binding. In vitro direct and competitive pull-down studies as well as western blot and RT-PCR analyses were used to validate predictions.
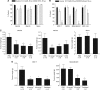
Results: The L1 loop (aa 112-124), a region within the S7-S8 loop (aa 214-236) and T140, P142, Q144, W146, R282 and L289 of the p53DBD were identified as potential sites for p28 binding. p28 decreased the level of the E3 ligase COP1 >80%, in p53wt and p53mut cells with no decrease in COP1 in p53dom/neg or p53null cells. Brief increases in the expression of the E3 ligases, TOPORS, Pirh2 and HDM2 (human double minute 2) in p53wt and p53mut cells were in response to sustained increases in p53.
Conclusion: These data identify the specific motifs within the DBD of p53 that bind p28 and suggest that p28 inhibition of COP1 binding results in the sustained, post-translational increase in p53 levels and subsequent inhibition of cancer cell growth independent of an HDM2 pathway.
Figures





Similar articles
-
A nanotechnological, molecular-modeling, and immunological approach to study the interaction of the anti-tumorigenic peptide p28 with the p53 family of proteins.Int J Nanomedicine. 2014 Apr 10;9:1799-813. doi: 10.2147/IJN.S58465. eCollection 2014. Int J Nanomedicine. 2014. PMID: 24748790 Free PMC article.
-
Interaction of an anticancer peptide fragment of azurin with p53 and its isolated domains studied by atomic force spectroscopy.Int J Nanomedicine. 2011;6:3011-9. doi: 10.2147/IJN.S26155. Epub 2011 Nov 24. Int J Nanomedicine. 2011. PMID: 22162658 Free PMC article.
-
Modelling the interaction between the p53 DNA-binding domain and the p28 peptide fragment of Azurin.J Mol Recognit. 2011 Nov-Dec;24(6):1043-55. doi: 10.1002/jmr.1153. J Mol Recognit. 2011. PMID: 22038811
-
p28 Bacterial Peptide, as an Anticancer Agent.Front Oncol. 2020 Aug 6;10:1303. doi: 10.3389/fonc.2020.01303. eCollection 2020. Front Oncol. 2020. PMID: 32850408 Free PMC article. Review.
-
Bacterial cupredoxin azurin hijacks cellular signaling networks: Protein-protein interactions and cancer therapy.Protein Sci. 2017 Dec;26(12):2334-2341. doi: 10.1002/pro.3310. Epub 2017 Oct 27. Protein Sci. 2017. PMID: 28960574 Free PMC article. Review.
Cited by
-
Dynamics of p53: A Master Decider of Cell Fate.Genes (Basel). 2017 Feb 9;8(2):66. doi: 10.3390/genes8020066. Genes (Basel). 2017. PMID: 28208785 Free PMC article. Review.
-
Bioactive peptides from food science to pharmaceutical industries: Their mechanism of action, potential role in cancer treatment and available resources.Heliyon. 2024 Nov 22;10(23):e40563. doi: 10.1016/j.heliyon.2024.e40563. eCollection 2024 Dec 15. Heliyon. 2024. PMID: 39654719 Free PMC article. Review.
-
Anticancer Actions of Azurin and Its Derived Peptide p28.Protein J. 2020 Apr;39(2):182-189. doi: 10.1007/s10930-020-09891-3. Protein J. 2020. PMID: 32180097 Review.
-
Regulation of the DNA damage response by ubiquitin conjugation.Front Genet. 2015 Mar 10;6:98. doi: 10.3389/fgene.2015.00098. eCollection 2015. Front Genet. 2015. PMID: 25806049 Free PMC article. Review.
-
The role of cell-penetrating peptides in potential anti-cancer therapy.Clin Transl Med. 2022 May;12(5):e822. doi: 10.1002/ctm2.822. Clin Transl Med. 2022. PMID: 35593206 Free PMC article. Review.
References
-
- Apiyo D, Wittung-Stafshede P. Unique complex between bacterial azurin and tumor-suppressor protein p53. Biochem Biophys Res Commun. 2005;332 (4:965–968. - PubMed
-
- Belmont LD, Mitchison TJ. Identification of a protein that interacts with tubulin dimers and increases the catastrophe rate of microtubules. Cell. 1996;84 (4:623–631. - PubMed
-
- Bianchi E, Denti S, Catena R, Rossetti G, Polo S, Gasparian S, Putignano S, Rogge L, Pardi R. Characterization of human constitutive photomorphogenesis protein 1, a RING finger ubiquitin ligase that interacts with Jun transcription factors and modulates their transcriptional activity. J Biol Chem. 2003;278 (22:19682–19690. - PubMed
-
- Bizzarri AR, Cannistraro S. Atomic force spectroscopy in biological complex formation: strategies and perspectives. J Phys Chem B. 2009;113 (52:16449–16464. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

