Network properties derived from deep sequencing of human B-cell receptor repertoires delineate B-cell populations
- PMID: 23742949
- PMCID: PMC3814887
- DOI: 10.1101/gr.154815.113
Network properties derived from deep sequencing of human B-cell receptor repertoires delineate B-cell populations
Abstract
The adaptive immune response selectively expands B- and T-cell clones following antigen recognition by B- and T-cell receptors (BCR and TCR), respectively. Next-generation sequencing is a powerful tool for dissecting the BCR and TCR populations at high resolution, but robust computational analyses are required to interpret such sequencing. Here, we develop a novel computational approach for BCR repertoire analysis using established next-generation sequencing methods coupled with network construction and population analysis. BCR sequences organize into networks based on sequence diversity, with differences in network connectivity clearly distinguishing between diverse repertoires of healthy individuals and clonally expanded repertoires from individuals with chronic lymphocytic leukemia (CLL) and other clonal blood disorders. Network population measures defined by the Gini Index and cluster sizes quantify the BCR clonality status and are robust to sampling and sequencing depths. BCR network analysis therefore allows the direct and quantifiable comparison of BCR repertoires between samples and intra-individual population changes between temporal or spatially separated samples and over the course of therapy.
Figures
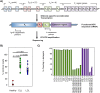



References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ 1990. Basic local alignment search tool. J Mol Biol 215: 403–410 - PubMed
-
- Bagnara D, Callea V, Stelitano C, Morabito F, Fabris S, Neri A, Zanardi S, Ghiotto F, Ciccone E, Grossi CE, et al. 2006. IgV gene intraclonal diversification and clonal evolution in B-cell chronic lymphocytic leukaemia. Br J Haematol 133: 50–58 - PubMed
-
- Batrak V, Blagodatski A, Buerstedde JM 2011. Understanding the immunoglobulin locus specificity of hypermutation. Methods Mol Biol 745: 311–326 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
