Ectopic overexpression of SsCBF1, a CRT/DRE-binding factor from the nightshade plant Solanum lycopersicoides, confers freezing and salt tolerance in transgenic Arabidopsis
- PMID: 23755095
- PMCID: PMC3670921
- DOI: 10.1371/journal.pone.0061810
Ectopic overexpression of SsCBF1, a CRT/DRE-binding factor from the nightshade plant Solanum lycopersicoides, confers freezing and salt tolerance in transgenic Arabidopsis
Abstract
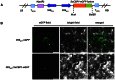
The C-repeat (CRT)/dehydration-responsive element (DRE) binding factor (CBF/DREB1) transcription factors play a key role in cold response. However, the detailed roles of many plant CBFs are far from fully understood. A CBF gene (SsCBF1) was isolated from the cold-hardy plant Solanum lycopersicoides. A subcellular localization study using GFP fusion protein indicated that SsCBF1 is localized in the nucleus. We delimited the SsCBF1 transcriptional activation domain to the C-terminal segment comprising amino acid residues 193-228 (SsCBF1(193-228)). The expression of SsCBF1 could be dramatically induced by cold, drought and high salinity. Transactivation assays in tobacco leaves revealed that SsCBF1 could specifically bind to the CRT cis-elements in vivo to activate the expression of downstream reporter genes. The ectopic overexpression of SsCBF1 conferred increased freezing and high-salinity tolerance and late flowering phenotype to transgenic Arabidopsis. RNA-sequencing data exhibited that a set of cold and salt stress responsive genes were up-regulated in transgenic Arabidopsis. Our results suggest that SsCBF1 behaves as a typical CBF to contribute to plant freezing tolerance. Increased resistance to high-salinity and late flowering phenotype derived from SsCBF1 OE lines lend more credence to the hypothesis that plant CBFs participate in diverse physiological and biochemical processes related to adverse conditions.
Conflict of interest statement
Figures







Similar articles
-
PpCBF3 from Cold-Tolerant Kentucky Bluegrass Involved in Freezing Tolerance Associated with Up-Regulation of Cold-Related Genes in Transgenic Arabidopsis thaliana.PLoS One. 2015 Jul 15;10(7):e0132928. doi: 10.1371/journal.pone.0132928. eCollection 2015. PLoS One. 2015. PMID: 26177510 Free PMC article.
-
Ectopic overexpression of SlHsfA3, a heat stress transcription factor from tomato, confers increased thermotolerance and salt hypersensitivity in germination in transgenic Arabidopsis.PLoS One. 2013;8(1):e54880. doi: 10.1371/journal.pone.0054880. Epub 2013 Jan 22. PLoS One. 2013. PMID: 23349984 Free PMC article.
-
Overexpression of a novel cold-responsive transcript factor LcFIN1 from sheepgrass enhances tolerance to low temperature stress in transgenic plants.Plant Biotechnol J. 2016 Mar;14(3):861-74. doi: 10.1111/pbi.12435. Epub 2015 Aug 3. Plant Biotechnol J. 2016. PMID: 26234381 Free PMC article.
-
[Arabidopsis CBF1 in plant tolerance to low temperature and drought stresses].Yi Chuan. 2004 May;26(3):394-8. Yi Chuan. 2004. PMID: 15640027 Review. Chinese.
-
Recent progress of salinity tolerance research in plants.Genetika. 2012 May;48(5):590-8. Genetika. 2012. PMID: 22830254 Review.
Cited by
-
Development of a PCR-based, genetic marker resource for the tomato-like nightshade relative, Solanum lycopersicoides using whole genome sequence analysis.PLoS One. 2020 Nov 23;15(11):e0242882. doi: 10.1371/journal.pone.0242882. eCollection 2020. PLoS One. 2020. PMID: 33227039 Free PMC article.
-
Molecular Evidence for Functional Divergence and Decay of a Transcription Factor Derived from Whole-Genome Duplication in Arabidopsis thaliana.Plant Physiol. 2015 Aug;168(4):1717-34. doi: 10.1104/pp.15.00689. Epub 2015 Jun 23. Plant Physiol. 2015. PMID: 26103993 Free PMC article.
-
Chilling acclimation provides immunity to stress by altering regulatory networks and inducing genes with protective functions in cassava.BMC Plant Biol. 2014 Aug 5;14:207. doi: 10.1186/s12870-014-0207-5. BMC Plant Biol. 2014. PMID: 25090992 Free PMC article.
-
Alien introgression and morpho-agronomic characterization of diploid progenies of Solanum lycopersicoides monosomic alien addition lines (MAALs) toward pre-breeding applications in tomato (S. lycopersicum).Theor Appl Genet. 2021 Apr;134(4):1133-1146. doi: 10.1007/s00122-020-03758-y. Epub 2021 Jan 2. Theor Appl Genet. 2021. PMID: 33386862 Free PMC article.
-
Phylogenomic analysis of cytochrome P450 multigene family and their differential expression analysis in Solanum lycopersicum L. suggested tissue specific promoters.BMC Genomics. 2019 Feb 7;20(1):116. doi: 10.1186/s12864-019-5483-x. BMC Genomics. 2019. PMID: 30732561 Free PMC article.
References
-
- Chinnusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends in Plant Science 12: 444–451. - PubMed
-
- Pino MT, Skinner JS, Jeknić Z, Hayes PM, Soeldner AH, et al. (2008) Ectopic AtCBF1 over-expression enhances freezing tolerance and induces cold acclimation-associated physiological modifications in potato. Plant, Cell & Environment 31: 393–406. - PubMed
-
- Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional Regulatory Networks in Cellular Responses and Tolerance to Dehydration and Cold Stresses. The Annual Review of Plant Biology 57: 781–803. - PubMed
-
- Jiang C, Iu B, Singh J (1996) Requirement of a CCGAC cis-acting element for cold induction of the BNll5 gene from winter Brassica napus. Plant Molecular Biology 30: 679–684. - PubMed
-
- Stockinger EJ, Gilmour SJ, Thomashow MF (1997) Arabidopsis thaliana CBF1 encodes an AP2 domain-containing transcriptional activator that binds to the C-repeatyDRE, a cis-acting DNA regulatory element that stimulates transcription in response to low temperature and water deficit. Proc Natl Acad Sci 94: 1035–1040. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

