Senescence-associated lncRNAs: senescence-associated long noncoding RNAs
- PMID: 23758631
- PMCID: PMC3773026
- DOI: 10.1111/acel.12115
Senescence-associated lncRNAs: senescence-associated long noncoding RNAs
Abstract
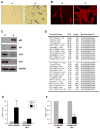
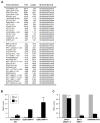
Noncoding RNAs include small transcripts, such as microRNAs and piwi-interacting RNAs, and a wide range of long noncoding RNAs (lncRNAs). Although many lncRNAs have been identified, only a small number of lncRNAs have been characterized functionally. Here, we sought to identify lncRNAs differentially expressed during replicative senescence. We compared lncRNAs expressed in proliferating, early-passage, 'young' human diploid WI-38 fibroblasts [population doubling (PDL) 20] with those expressed in senescent, late-passage, 'old' fibroblasts (PDL 52) by RNA sequencing (RNA-Seq). Numerous transcripts in all lncRNA groups (antisense lncRNAs, pseudogene-encoded lncRNAs, previously described lncRNAs and novel lncRNAs) were validated using reverse transcription (RT) and real-time, quantitative (q)PCR. Among the novel senescence-associated lncRNAs (SAL-RNAs) showing lower abundance in senescent cells, SAL-RNA1 (XLOC_023166) was found to delay senescence, because reducing SAL-RNA1 levels enhanced the appearance of phenotypic traits of senescence, including an enlarged morphology, positive β-galactosidase activity, and heightened p53 levels. Our results reveal that the expression of known and novel lncRNAs changes with senescence and suggests that SAL-RNAs play direct regulatory roles in this important cellular process.
Keywords: noncoding; post-transcriptional gene regulation; proliferation; senescence-associated gene expression patterns; transcriptome.
Published 2013. This article is a U.S. Government work and is in the public domain in the USA.
Conflict of interest statement
The authors do not have any conflicts of interest.
Figures




References
-
- Campisi J. Senescent cells, tumor suppression, and organismal aging: good citizens, bad neighbors. Cell. 2005;120:513–522. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

