Investigation of dmyc Promoter and Regulatory Regions
- PMID: 23761963
- PMCID: PMC3663572
- DOI: 10.4137/GRSB.S10751
Investigation of dmyc Promoter and Regulatory Regions
Abstract
Products of the myc gene family integrate extracellular signals by modulating a wide range of their targets involved in cellular biogenesis and metabolism; the purpose of this integration is to regulate cell death, proliferation, and differentiation. However, understanding the regulation of myc at the transcription level remains a challenge. We performed rapid amplification of dmyc cDNA ends (5' RACE) and mapped the transcription start site at P1 promoter, 18 base pairs upstream of the start of the known EST GM01143 and within the 5' UTR. Our data show that the first TATA box, previously computationally predicted, is utilized to generate dmyc full length mRNA. The largest transcript contains all three exons, generated after the removal of the introns by constitutively regulated splicing events. Further investigation of Downstream Promoter Element (DPE) was achieved by studying lacZ reporter activity; investigation revealed that this element and its upstream cluster of binding sites are required for the dmyc intron 2 activity. These findings may provide valuable tools for further analysis of dmyc cis-elements.
Keywords: 5′ RACE; Downstream Promoter Element (DPE); Drosophila; RNA splicing; TATA-box; dmyc.
Figures


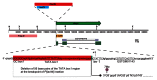





References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

