Global expression profiling of transcription factor genes provides new insights into pathogenicity and stress responses in the rice blast fungus
- PMID: 23762023
- PMCID: PMC3675110
- DOI: 10.1371/journal.ppat.1003350
Global expression profiling of transcription factor genes provides new insights into pathogenicity and stress responses in the rice blast fungus
Abstract
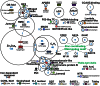
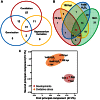
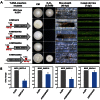
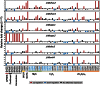
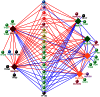
Because most efforts to understand the molecular mechanisms underpinning fungal pathogenicity have focused on studying the function and role of individual genes, relatively little is known about how transcriptional machineries globally regulate and coordinate the expression of a large group of genes involved in pathogenesis. Using quantitative real-time PCR, we analyzed the expression patterns of 206 transcription factor (TF) genes in the rice blast fungus Magnaporthe oryzae under 32 conditions, including multiple infection-related developmental stages and various abiotic stresses. The resulting data, which are publicly available via an online platform, provided new insights into how these TFs are regulated and potentially work together to control cellular responses to a diverse array of stimuli. High degrees of differential TF expression were observed under the conditions tested. More than 50% of the 206 TF genes were up-regulated during conidiation and/or in conidia. Mutations in ten conidiation-specific TF genes caused defects in conidiation. Expression patterns in planta were similar to those under oxidative stress conditions. Mutants of in planta inducible genes not only exhibited sensitive to oxidative stress but also failed to infect rice. These experimental validations clearly demonstrated the value of TF expression patterns in predicting the function of individual TF genes. The regulatory network of TF genes revealed by this study provides a solid foundation for elucidating how M. oryzae regulates its pathogenesis, development, and stress responses.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Systematic analysis of Zn2Cys6 transcription factors required for development and pathogenicity by high-throughput gene knockout in the rice blast fungus.PLoS Pathog. 2014 Oct 9;10(10):e1004432. doi: 10.1371/journal.ppat.1004432. eCollection 2014 Oct. PLoS Pathog. 2014. PMID: 25299517 Free PMC article.
-
Characterization of 47 Cys2 -His2 zinc finger proteins required for the development and pathogenicity of the rice blast fungus Magnaporthe oryzae.New Phytol. 2016 Aug;211(3):1035-51. doi: 10.1111/nph.13948. Epub 2016 Apr 4. New Phytol. 2016. PMID: 27041000
-
MoSnt2-dependent deacetylation of histone H3 mediates MoTor-dependent autophagy and plant infection by the rice blast fungus Magnaporthe oryzae.Autophagy. 2018;14(9):1543-1561. doi: 10.1080/15548627.2018.1458171. Epub 2018 Aug 31. Autophagy. 2018. PMID: 29929416 Free PMC article.
-
Two conidiation-related Zn(II)2Cys6 transcription factor genes in the rice blast fungus.Fungal Genet Biol. 2013 Dec;61:133-41. doi: 10.1016/j.fgb.2013.10.004. Epub 2013 Oct 15. Fungal Genet Biol. 2013. PMID: 24140150
-
Roles of Peroxisomes in the Rice Blast Fungus.Biomed Res Int. 2016;2016:9343417. doi: 10.1155/2016/9343417. Epub 2016 Aug 16. Biomed Res Int. 2016. PMID: 27610388 Free PMC article. Review.
Cited by
-
Genome-Wide Identification and Expression Analysis of the Basic Leucine Zipper (bZIP) Transcription Factor Gene Family in Fusarium graminearum.Genes (Basel). 2022 Mar 28;13(4):607. doi: 10.3390/genes13040607. Genes (Basel). 2022. PMID: 35456413 Free PMC article.
-
NADPH Oxidases Are Required for Appressorium-Mediated Penetration in Colletotrichum scovillei-Pepper Fruit Pathosystem.Plant Pathol J. 2022 Aug;38(4):345-354. doi: 10.5423/PPJ.OA.05.2022.0066. Epub 2022 Aug 1. Plant Pathol J. 2022. PMID: 35953054 Free PMC article.
-
Transcriptome profiling reveals insertional mutagenesis suppressed the expression of candidate pathogenicity genes in honeybee fungal pathogen, Ascosphaera apis.Sci Rep. 2020 May 5;10(1):7532. doi: 10.1038/s41598-020-64022-3. Sci Rep. 2020. PMID: 32372055 Free PMC article.
-
Roles of Forkhead-box Transcription Factors in Controlling Development, Pathogenicity, and Stress Response in Magnaporthe oryzae.Plant Pathol J. 2014 Jun;30(2):136-50. doi: 10.5423/PPJ.OA.02.2014.0018. Plant Pathol J. 2014. PMID: 25288996 Free PMC article.
-
Comparative profiling of canonical and non-canonical small RNAs in the rice blast fungus, Magnaporthe oryzae.Front Microbiol. 2022 Sep 26;13:995334. doi: 10.3389/fmicb.2022.995334. eCollection 2022. Front Microbiol. 2022. PMID: 36225371 Free PMC article.
References
-
- Riechmann JL, Heard J, Martin G, Reuber L, Jiang C, et al. (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290: 2105–2110. - PubMed
-
- Czechowski T, Bari RP, Stitt M, Scheible WR, Udvardi MK (2004) Real-time RT-PCR profiling of over 1400 Arabidopsis transcription factors: unprecedented sensitivity reveals novel root- and shoot-specific genes. Plant J 38: 366–379. - PubMed
-
- Belluardo N, Olsson PA, Mudo G, Sommer WH, Amato G, et al. (2005) Transcription factor gene expression profiling after acute intermittent nicotine treatment in the rat cerebral cortex. Neuroscience 133: 787–796. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

