Genome-wide genetic diversity and differentially selected regions among Suffolk, Rambouillet, Columbia, Polypay, and Targhee sheep
- PMID: 23762451
- PMCID: PMC3677876
- DOI: 10.1371/journal.pone.0065942
Genome-wide genetic diversity and differentially selected regions among Suffolk, Rambouillet, Columbia, Polypay, and Targhee sheep
Abstract
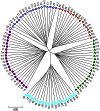
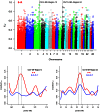
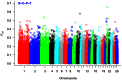
Sheep are among the major economically important livestock species worldwide because the animals produce milk, wool, skin, and meat. In the present study, the Illumina OvineSNP50 BeadChip was used to investigate genetic diversity and genome selection among Suffolk, Rambouillet, Columbia, Polypay, and Targhee sheep breeds from the United States. After quality-control filtering of SNPs (single nucleotide polymorphisms), we used 48,026 SNPs, including 46,850 SNPs on autosomes that were in Hardy-Weinberg equilibrium and 1,176 SNPs on chromosome × for analysis. Phylogenetic analysis based on all 46,850 SNPs clearly separated Suffolk from Rambouillet, Columbia, Polypay, and Targhee, which was not surprising as Rambouillet contributed to the synthesis of the later three breeds. Based on pair-wise estimates of F(ST), significant genetic differentiation appeared between Suffolk and Rambouillet (F(ST) = 0.1621), while Rambouillet and Targhee had the closest relationship (F(ST) = 0.0681). A scan of the genome revealed 45 and 41 differentially selected regions (DSRs) between Suffolk and Rambouillet and among Rambouillet-related breed populations, respectively. Our data indicated that regions 13 and 24 between Suffolk and Rambouillet might be good candidates for evaluating breed differences. Furthermore, ovine genome v3.1 assembly was used as reference to link functionally known homologous genes to economically important traits covered by these differentially selected regions. In brief, our present study provides a comprehensive genome-wide view on within- and between-breed genetic differentiation, biodiversity, and evolution among Suffolk, Rambouillet, Columbia, Polypay, and Targhee sheep breeds. These results may provide new guidance for the synthesis of new breeds with different breeding objectives.
Conflict of interest statement
Figures

Similar articles
-
Genetic relationship between milk score and litter weight for Targhee, Columbia, Rambouillet, and Polypay sheep.J Anim Sci. 2005 Apr;83(4):786-93. doi: 10.2527/2005.834786x. J Anim Sci. 2005. PMID: 15753332
-
Usefulness of subjective ovine milk scores: II. Genetic parameter estimates.J Anim Sci. 2001 Apr;79(4):869-76. doi: 10.2527/2001.794869x. J Anim Sci. 2001. PMID: 11325191
-
Influence of body weight, age, and weight gain on fertility and prolificacy in four breeds of ewe lambs.J Anim Sci. 2005 Jul;83(7):1680-9. doi: 10.2527/2005.8371680x. J Anim Sci. 2005. PMID: 15956477
-
Whole-Genome Resequencing in Sheep: Applications in Breeding, Evolution, and Conservation.Genes (Basel). 2025 Mar 22;16(4):363. doi: 10.3390/genes16040363. Genes (Basel). 2025. PMID: 40282323 Free PMC article. Review.
-
Genetics of the phenotypic evolution in sheep: a molecular look at diversity-driving genes.Genet Sel Evol. 2022 Sep 9;54(1):61. doi: 10.1186/s12711-022-00753-3. Genet Sel Evol. 2022. PMID: 36085023 Free PMC article. Review.
Cited by
-
A high-density genome-wide association with absolute blood monocyte count in domestic sheep identifies novel loci.PLoS One. 2022 May 6;17(5):e0266748. doi: 10.1371/journal.pone.0266748. eCollection 2022. PLoS One. 2022. PMID: 35522671 Free PMC article.
-
An improved ovine reference genome assembly to facilitate in-depth functional annotation of the sheep genome.Gigascience. 2022 Feb 4;11:giab096. doi: 10.1093/gigascience/giab096. Gigascience. 2022. PMID: 35134925 Free PMC article.
-
Genome-Wide Specific Selection in Three Domestic Sheep Breeds.PLoS One. 2015 Jun 17;10(6):e0128688. doi: 10.1371/journal.pone.0128688. eCollection 2015. PLoS One. 2015. PMID: 26083354 Free PMC article.
-
A Combined Multi-Cohort Approach Reveals Novel and Known Genome-Wide Selection Signatures for Wool Traits in Merino and Merino-Derived Sheep Breeds.Front Genet. 2019 Oct 25;10:1025. doi: 10.3389/fgene.2019.01025. eCollection 2019. Front Genet. 2019. PMID: 31708969 Free PMC article.
-
Genomic Population Structure of the Main Historical Genetic Lines of Spanish Merino Sheep.Animals (Basel). 2022 May 23;12(10):1327. doi: 10.3390/ani12101327. Animals (Basel). 2022. PMID: 35625173 Free PMC article.
References
-
- Goldammer T, Di Meo GP, Lühken G, Drögemüller C, Wu CH, et al. (2009) Molecular cytogenetics and gene mapping in sheep (Ovis aries, 2n = 54). Cytogenet Genome Res126: 63–76. - PubMed
-
- International Sheep Genomics Consortium, Archibald AL, Cockett NE, Dalrymple BP, Faraut T, et al. (2010) The sheep genome reference sequence: a work in progress. Anim Genet 41: 449–453. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

