Revealing the hidden relationship by sparse modules in complex networks with a large-scale analysis
- PMID: 23762457
- PMCID: PMC3677904
- DOI: 10.1371/journal.pone.0066020
Revealing the hidden relationship by sparse modules in complex networks with a large-scale analysis
Abstract
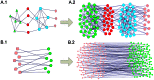
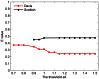
One of the remarkable features of networks is module that can provide useful insights into not only network organizations but also functional behaviors between their components. Comprehensive efforts have been devoted to investigating cohesive modules in the past decade. However, it is still not clear whether there are important structural characteristics of the nodes that do not belong to any cohesive module. In order to answer this question, we performed a large-scale analysis on 25 complex networks with different types and scales using our recently developed BTS (bintree seeking) algorithm, which is able to detect both cohesive and sparse modules in the network. Our results reveal that the sparse modules composed by the cohesively isolated nodes widely co-exist with the cohesive modules. Detailed analysis shows that both types of modules provide better characterization for the division of a network into functional units than merely cohesive modules, because the sparse modules possibly re-organize the nodes in the so-called cohesive modules, which lack obvious modular significance, into meaningful groups. Compared with cohesive modules, the sizes of sparse ones are generally smaller. Sparse modules are also found to have preferences in social and biological networks than others.
Conflict of interest statement
Figures
Similar articles
-
BinTree seeking: a novel approach to mine both bi-sparse and cohesive modules in protein interaction networks.PLoS One. 2011;6(11):e27646. doi: 10.1371/journal.pone.0027646. Epub 2011 Nov 28. PLoS One. 2011. PMID: 22140454 Free PMC article.
-
A new multi-scale method to reveal hierarchical modular structures in biological networks.Mol Biosyst. 2016 Nov 15;12(12):3724-3733. doi: 10.1039/c6mb00617e. Mol Biosyst. 2016. PMID: 27783080
-
Protein interaction networks--more than mere modules.PLoS Comput Biol. 2010 Jan 29;6(1):e1000659. doi: 10.1371/journal.pcbi.1000659. PLoS Comput Biol. 2010. PMID: 20126533 Free PMC article.
-
Identifying protein complexes and functional modules--from static PPI networks to dynamic PPI networks.Brief Bioinform. 2014 Mar;15(2):177-94. doi: 10.1093/bib/bbt039. Epub 2013 Jun 18. Brief Bioinform. 2014. PMID: 23780996 Review.
-
Learning Differential Module Networks Across Multiple Experimental Conditions.Methods Mol Biol. 2019;1883:303-321. doi: 10.1007/978-1-4939-8882-2_13. Methods Mol Biol. 2019. PMID: 30547406 Review.
Cited by
-
Matching rules for collective behaviors on complex networks: optimal configurations for vibration frequencies of networked harmonic oscillators.PLoS One. 2013 Dec 26;8(12):e82161. doi: 10.1371/journal.pone.0082161. eCollection 2013. PLoS One. 2013. PMID: 24386088 Free PMC article.
References
-
- Newman MEJ, Girvan M (2004) Finding and evaluating community structure in networks. Physical Review E 69: 026113. - PubMed
-
- Flake GW, Lawrence S, Giles CL, Coetzee FM (2002) Self-organization and identification of web communities. Computer 35: 66–70.
-
- Barabasi AL, Oltvai ZN (2004) Network biology: Understanding the cell’s functional organization. Nature Reviews Genetics 5: 101–U115. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

