Combined methylation mapping of 5mC and 5hmC during early embryonic stages in bovine
- PMID: 23773395
- PMCID: PMC3689598
- DOI: 10.1186/1471-2164-14-406
Combined methylation mapping of 5mC and 5hmC during early embryonic stages in bovine
Abstract
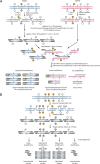
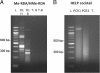
Background: It was recently established that changes in methylation during development are dynamic and involve both methylation and demethylation processes. Yet, which genomic sites are changing and what are the contributions of methylation (5mC) and hydroxymethylation (5hmC) to this epigenetic remodeling is still unknown. When studying early development, options for methylation profiling are limited by the unavailability of sufficient DNA material from these scarce samples and limitations are aggravated in non-model species due to the lack of technological platforms. We therefore sought to obtain a representation of differentially 5mC or 5hmC loci during bovine early embryo stages through the use of three complementary methods, based on selective methyl-sensitive restriction and enrichment by ligation-mediated PCR or on subtractive hybridization. Using these strategies, libraries of putative methylation and hydroxymethylated sites were generated from Day-7 and Day-12 bovine embryos.
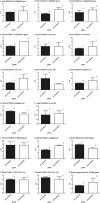
Results: Over 1.2 million sequencing reads were analyzed, resulting in 151,501 contigs, of which 69,136 were uniquely positioned on the genome. A total of 101,461 putative methylated sites were identified. The output of the three methods differed in genomic coverage as well as in the nature of the identified sites. The classical MspI/HpaII combination of restriction enzymes targeted CpG islands whereas the other methods covered 5mC and 5hmC sites outside of these regions. Data analysis suggests a transition of these methylation marks between Day-7 and Day-12 embryos in specific classes of repeat-containing elements.
Conclusions: Our combined strategy offers a genomic map of the distribution of cytosine methylation/hydroxymethylation during early bovine embryo development. These results support the hypothesis of a regulatory phase of hypomethylation in repeat sequences during early embryogenesis.
Figures



References
-
- Ehrlich M, Buchanan KL, Tsien F, Jiang G, Sun B, Uicker W, Weemaes CMR, Smeets D, Sperling K, Belohradsky BH. et al. DNA methyltransferase 3B mutations linked to the ICF syndrome cause dysregulation of lymphogenesis genes. Hum Mol Genet. 2001;10(25):2917–2931. doi: 10.1093/hmg/10.25.2917. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

