Dissecting a complex chemical stress: chemogenomic profiling of plant hydrolysates
- PMID: 23774757
- PMCID: PMC3964314
- DOI: 10.1038/msb.2013.30
Dissecting a complex chemical stress: chemogenomic profiling of plant hydrolysates
Abstract
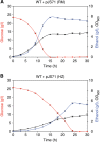
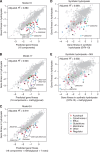
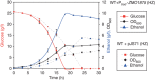
The efficient production of biofuels from cellulosic feedstocks will require the efficient fermentation of the sugars in hydrolyzed plant material. Unfortunately, plant hydrolysates also contain many compounds that inhibit microbial growth and fermentation. We used DNA-barcoded mutant libraries to identify genes that are important for hydrolysate tolerance in both Zymomonas mobilis (44 genes) and Saccharomyces cerevisiae (99 genes). Overexpression of a Z. mobilis tolerance gene of unknown function (ZMO1875) improved its specific ethanol productivity 2.4-fold in the presence of miscanthus hydrolysate. However, a mixture of 37 hydrolysate-derived inhibitors was not sufficient to explain the fitness profile of plant hydrolysate. To deconstruct the fitness profile of hydrolysate, we profiled the 37 inhibitors against a library of Z. mobilis mutants and we modeled fitness in hydrolysate as a mixture of fitness in its components. By examining outliers in this model, we identified methylglyoxal as a previously unknown component of hydrolysate. Our work provides a general strategy to dissect how microbes respond to a complex chemical stress and should enable further engineering of hydrolysate tolerance.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


References
-
- Agrawal M, Wang Y, Chen RR (2012) Engineering efficient xylose metabolism into an acetic acid-tolerant Zymomonas mobilis strain by introducing adaptation-induced mutations. Biotechnol Lett 34: 1825–1832 - PubMed
-
- Almeida JR, Modig T, Petersson A, Hahn-Hagerdal B, Lidén G, Gorwa-Grauslund M-F (2007) Increased tolerance and conversion of inhibitors in lignocellulosic hydrolysates by Saccharomyces cerevisiae. J Chem Technol Biotechnol 82: 340–349
-
- Amin G, Van den Eynde E, Verachtert H (1983) Determination of by-products formed during the ethanolic fermentation, using batch and immobilized cell systems of Zymomonas mobilis and Saccharomyces bayanus. Eur J Appl Microbiol Biotechnol 18: 1–5
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

